 Back
BackThe Molecular Basis of Heredity: DNA Structure and Replication
Study Guide - Smart Notes

Macromolecules and Their Role in Genetics
Overview of Biological Macromolecules
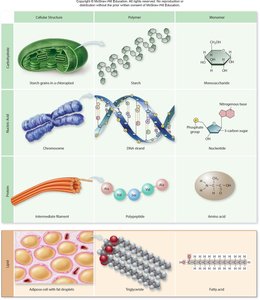
Macromolecules are large, complex molecules essential for life, including carbohydrates, lipids, proteins, and nucleic acids. Each class is composed of specific monomers that polymerize to form functional polymers, which play critical roles in cellular structure and function.
Carbohydrates: Monomers are monosaccharides; polymers include starch and cellulose.
Lipids: Not true polymers, but composed of fatty acids and glycerol; form structures like triglycerides.
Proteins: Monomers are amino acids; polymers are polypeptides and functional proteins.
Nucleic Acids: Monomers are nucleotides; polymers are DNA and RNA.
Polymer Assembly and Disassembly
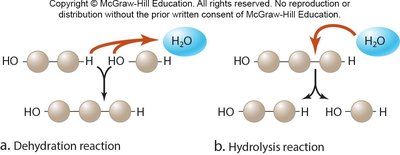
Polymers are assembled and disassembled through specific chemical reactions:
Dehydration Synthesis: Joins monomers by removing a water molecule, forming a covalent bond.
Hydrolysis: Breaks polymers into monomers by adding water, cleaving the covalent bond.
Functional Groups in Macromolecules
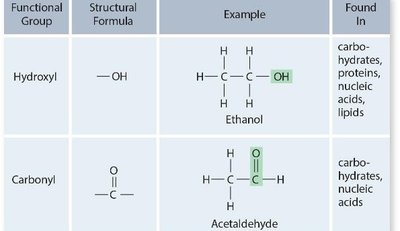
Functional groups are specific groups of atoms within molecules that determine the chemical properties and reactivity of those molecules. They are critical in the structure and function of all macromolecules.
Hydroxyl (-OH): Found in carbohydrates, proteins, nucleic acids, and lipids.
Carbonyl (C=O): Present in carbohydrates and nucleic acids.
Other groups include amino, carboxyl, phosphate, and sulfhydryl.
Nucleic Acids: Structure and Function
Nucleotides: The Building Blocks of Nucleic Acids
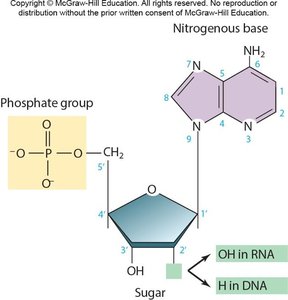
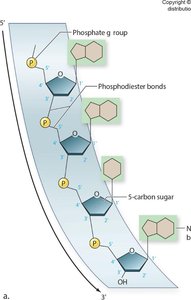
Nucleotides are the monomers of nucleic acids, each consisting of a five-carbon sugar, a phosphate group, and a nitrogenous base. The sugar is either ribose (in RNA) or deoxyribose (in DNA).
Nitrogenous Bases: Purines (adenine, guanine) and pyrimidines (cytosine, thymine in DNA; uracil in RNA).
Phosphate Group: Links nucleotides together via phosphodiester bonds.
Structure of Nucleic Acid Polymers
Nucleic acids are linear polymers of nucleotides connected by phosphodiester bonds, forming a sugar-phosphate backbone with protruding nitrogenous bases.
Directionality: Nucleic acids have a 5' to 3' orientation, critical for replication and transcription.
DNA and RNA: Structural Comparison
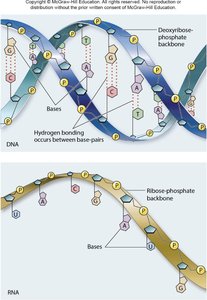
DNA and RNA differ in their sugar component, nitrogenous bases, and overall structure:
DNA: Double-stranded, deoxyribose sugar, bases A, T, G, C.
RNA: Single-stranded, ribose sugar, bases A, U, G, C.
Proteins: Structure and Relevance to Genetics
Amino Acids and Protein Structure
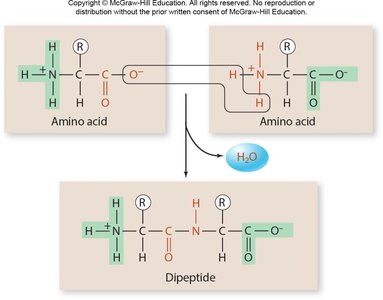
Proteins are polymers of amino acids, each with a central carbon atom bonded to an amino group, a carboxyl group, a hydrogen atom, and a variable R group. The sequence and chemical properties of R groups determine protein structure and function.
Peptide Bonds: Link amino acids via dehydration synthesis, forming polypeptides.
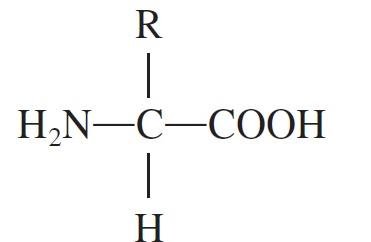
The Molecular Basis of Heredity: DNA as Genetic Material
Experimental Evidence for DNA as Genetic Material
Multiple experiments established DNA as the hereditary material:
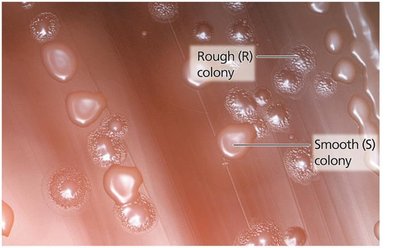
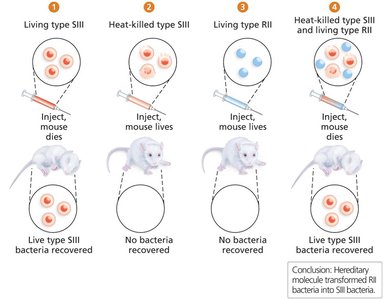
Griffith's Transformation Experiment (1928): Showed that a 'transforming factor' from dead virulent bacteria could genetically alter non-virulent bacteria.
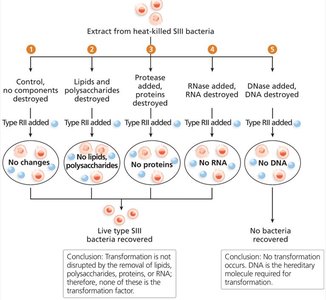
Avery, MacLeod, and McCarty (1944): Identified DNA as the transforming factor by selectively destroying proteins, RNA, or DNA.
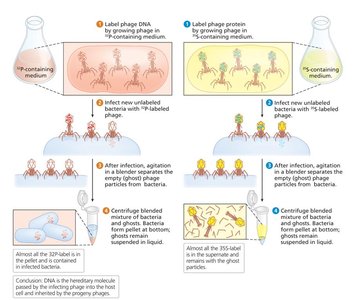
Hershey-Chase Experiment (1952): Used bacteriophages labeled with radioactive isotopes to confirm DNA, not protein, is the genetic material.
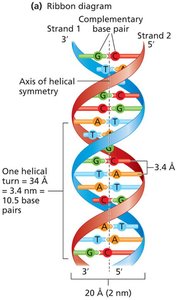
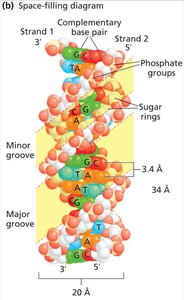
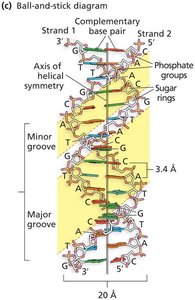
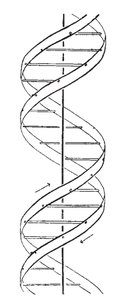
DNA Structure: The Double Helix
The structure of DNA was elucidated through several key findings:
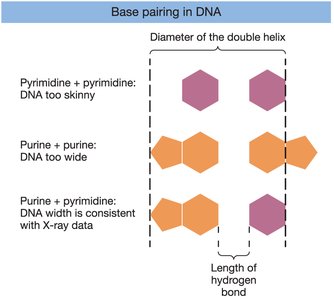
Chargaff's Rules: The amount of adenine equals thymine, and guanine equals cytosine (A=T, G=C).
Rosalind Franklin's X-ray Diffraction: Revealed the helical structure of DNA.
Watson and Crick Model (1953): Proposed the double helix with antiparallel strands and complementary base pairing.


DNA Replication: Mechanism and Enzymes
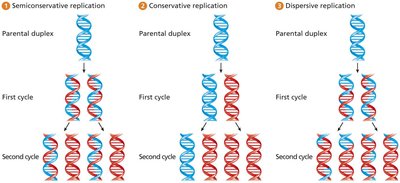
Semiconservative Replication
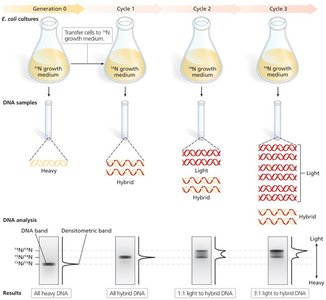
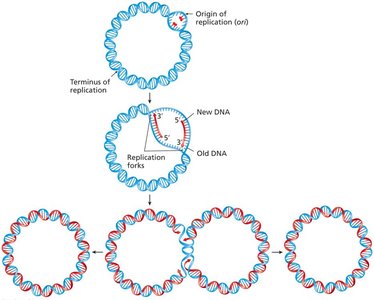
DNA replication is semiconservative, meaning each daughter DNA molecule consists of one parental and one newly synthesized strand. This was demonstrated by the Meselson-Stahl experiment (1958).
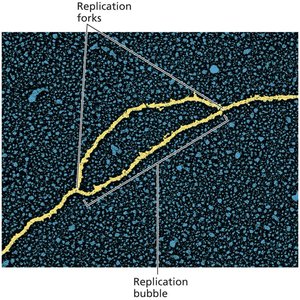
Replication Fork: The site where DNA unwinds and new strands are synthesized.
Bidirectional Replication: Replication proceeds in both directions from the origin.
Enzymes and Proteins in DNA Replication
DNA replication requires a coordinated set of enzymes and proteins:
Helicase: Unwinds the DNA double helix.
Single-Stranded Binding Proteins (SSBPs): Stabilize unwound DNA.
Primase: Synthesizes RNA primers.
DNA Polymerase III: Synthesizes new DNA strands.
DNA Polymerase I: Removes RNA primers and fills in with DNA.
Ligase: Joins Okazaki fragments on the lagging strand.
Topoisomerase (Gyrase): Relieves supercoiling ahead of the replication fork.
Initiation and Termination of Replication
Replication begins at specific origins of replication and ends with the completion of new DNA molecules. In eukaryotes, multiple origins exist per chromosome, while prokaryotes typically have a single origin.
Telomeres and Telomerase: In eukaryotes, telomerase extends chromosome ends to prevent loss of genetic material during replication.
Accuracy and Proofreading
DNA replication is highly accurate due to the proofreading activity of DNA polymerases, which correct errors during synthesis, resulting in a very low mutation rate.
Summary Table: Key Enzymes in DNA Replication
Function | Bacteria (E. coli) | Eukaryotes (Humans) |
|---|---|---|
Recognizes origins | DnaA | ORC (origin recognition complex) |
Unwinds double-stranded DNA | DnaB helicase | MCM2-7 helicase |
Assists helicase binding | DnaC | Cdc6 and ORC |
Stabilizes single-stranded DNA | SSB | RPA |
Removes supercoils | Gyrase | Topoisomerases |
Synthesizes RNA primers | Primase | DNA pol α-primase complex |
Elongates DNA | DNA pol III | DNA pol ε (leading), δ (lagging) |
Sliding clamp | β-clamp | PCNA |
Clamp loader | τ complex | RFC |
Removes RNA primers | DNA pol I | FEN1 |
Replaces RNA primers with DNA | DNA pol I | DNA pol ε |
Ligates Okazaki fragments | DNA ligase | DNA ligase I |
Additional info: This guide covers foundational concepts in molecular genetics, including the chemical nature of hereditary material, the structure of DNA, and the mechanisms of DNA replication. Understanding these principles is essential for advanced study in genetics, molecular biology, and related biomedical sciences.
