 Back
BackChapter 9
Study Guide - Smart Notes

The Molecular Biology of Translation
Overview of Genetic Information Flow
Translation is the process by which the genetic code carried by mRNA is decoded to produce a specific polypeptide, or protein. This process is a key component of the central dogma of molecular biology, which describes the flow of genetic information from DNA to RNA to protein.
Transcription: DNA is transcribed into messenger RNA (mRNA).
Translation: mRNA is translated into a polypeptide chain at the ribosome.
Polypeptide directionality: The N-terminus (amino end) corresponds to the start codon, and the C-terminus (carboxyl end) corresponds to the stop codon.
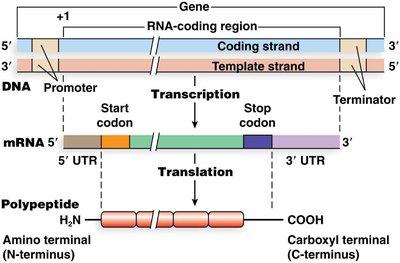
mRNA Structure and Codons
Messenger RNA (mRNA) contains both coding and non-coding regions that are essential for translation.
5' and 3' Untranslated Regions (UTRs): Segments outside the translated region; contain regulatory sequences such as the Shine-Dalgarno (bacteria) or Kozak (eukaryotes) sequences.
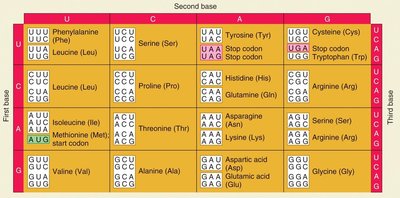
Codons: Triplets of nucleotides (A, U, C, G) that specify amino acids. There are 64 possible codons for 20 standard amino acids, making the code redundant but not ambiguous.
Start Codon: AUG (codes for methionine).
Stop Codons: UAA, UAG, UGA (do not code for any amino acid).
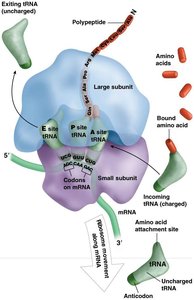
tRNA and Ribosome Structure
Transfer RNA (tRNA) and ribosomes are essential for decoding mRNA and synthesizing proteins.
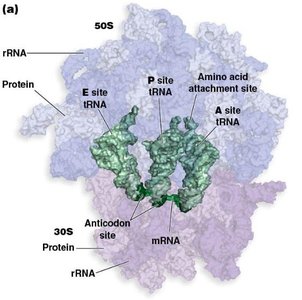
tRNA: Adaptor molecules with an anticodon region that base-pairs with mRNA codons and an acceptor stem for amino acid attachment.
Charging: Aminoacyl-tRNA synthetases attach the correct amino acid to each tRNA using ATP.
Wobble Hypothesis: Flexible base pairing at the third codon position allows fewer tRNA types to recognize multiple codons.
Ribosome: Composed of large and small subunits, with three key sites: A (aminoacyl), P (peptidyl), and E (exit).
Stages of Translation
Translation occurs in three main stages: initiation, elongation, and termination.
Initiation
Bacteria: The small ribosomal subunit binds the Shine-Dalgarno sequence on mRNA, and the initiator tRNA (fMet-tRNAfMet) pairs with the start codon.
Eukaryotes: The small subunit, initiation factors, and initiator tRNA scan the mRNA for the Kozak sequence and start codon.
Large ribosomal subunit joins, forming the complete initiation complex.
Elongation
Charged tRNAs are recruited to the A site.
Peptidyl transferase catalyzes peptide bond formation between amino acids at the P and A sites.
The ribosome translocates along the mRNA, moving the tRNA from the A site to the P site, and the uncharged tRNA exits via the E site.

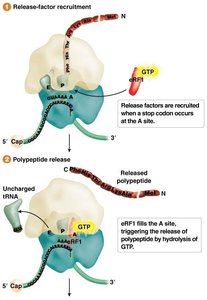
Termination
When a stop codon enters the A site, release factors bind and trigger the release of the polypeptide chain.
The ribosome dissociates, and the mRNA is released.
Comparing Bacterial and Eukaryotic Translation
There are key differences between translation in prokaryotes and eukaryotes:
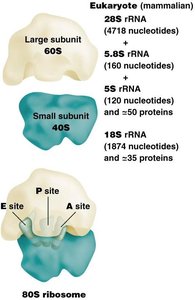
Ribosome Size: Prokaryotes have 70S ribosomes (50S + 30S), eukaryotes have 80S ribosomes (60S + 40S).
Initiation Sequences: Shine-Dalgarno in bacteria, Kozak in eukaryotes.
Simultaneity: In bacteria, transcription and translation can occur simultaneously; in eukaryotes, they are separated by the nuclear envelope.
Protein Structure and Function
The sequence of amino acids in a polypeptide determines its three-dimensional structure and function. The side chains (R groups) of amino acids confer specific chemical properties, influencing protein folding and activity.
Primary Structure: Linear sequence of amino acids.
Secondary, Tertiary, and Quaternary Structures: Higher-order folding and assembly of polypeptide chains.
Mutations: Single nucleotide changes can have significant or negligible effects on protein function, depending on their location and nature.
