 Back
BackThe Molecular Biology of Translation: From Genetic Code to Protein Synthesis
Study Guide - Smart Notes

The Molecular Biology of Translation
Introduction to Translation
Translation is the process by which the genetic information encoded in messenger RNA (mRNA) is used to synthesize proteins. This process is fundamental to gene expression and is a key step in the central dogma of molecular biology, which describes the flow of genetic information from DNA to RNA to protein.
Central Dogma: DNA is transcribed into RNA, which is then translated into protein.
Translation: Occurs in the cytoplasm at the ribosome, where mRNA codons are read and translated into a specific sequence of amino acids.
Key Players: mRNA, transfer RNA (tRNA), ribosomes, and various protein factors.
Flow of Genetic Information
From DNA to Protein
The genetic code is transcribed from DNA to mRNA and then translated into a polypeptide chain. The boundaries of translation are defined by the start and stop codons on the mRNA.
Start Codon (AUG): Specifies methionine and marks the beginning of translation.
Stop Codons (UAA, UAG, UGA): Signal the end of translation.
Untranslated Regions (UTRs): 5' and 3' UTRs are important for regulation but are not translated into protein.
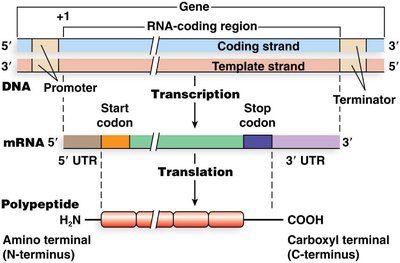
mRNA and the Genetic Code
Codons and the Genetic Code
The genetic code consists of triplet codons, each specifying a particular amino acid. The code is nearly universal, non-overlapping, and degenerate (redundant).
Codon: A sequence of three nucleotides on mRNA that codes for a specific amino acid.
Degeneracy: Multiple codons can code for the same amino acid.
Universality: The genetic code is conserved across most organisms, with minor exceptions.
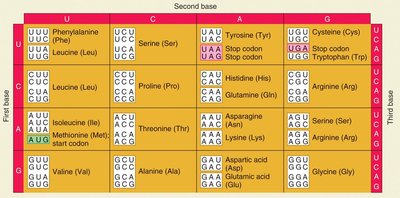
tRNA Structure and Function
tRNA: The Adapter Molecule
Transfer RNA (tRNA) molecules serve as adapters that match specific codons in the mRNA with their corresponding amino acids during translation. Each tRNA has an anticodon region that base-pairs with the mRNA codon and an acceptor stem for amino acid attachment.
Anticodon: A sequence of three bases on tRNA complementary to the mRNA codon.
Amino Acid Attachment Site: The 3' end of tRNA where the amino acid is covalently attached.
Modified Bases: tRNAs contain unusual bases (e.g., inosine) that contribute to their structure and function.
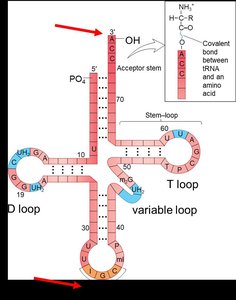
Charging tRNA: Aminoacyl-tRNA Synthetase
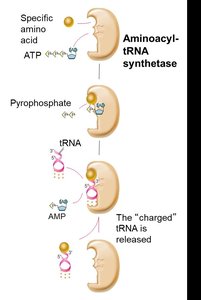
tRNAs are charged with their respective amino acids by specific enzymes called aminoacyl-tRNA synthetases. This process is essential for translation fidelity and requires energy from ATP hydrolysis.
Step 1: Amino acid activation by ATP, forming aminoacyl-AMP.
Step 2: Transfer of the amino acid to the tRNA, forming aminoacyl-tRNA (charged tRNA).
Specificity: Each amino acid has its own synthetase, ensuring correct pairing.
Wobble Hypothesis
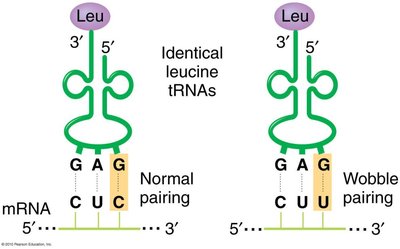
The number of tRNA types is less than the number of codons due to the wobble base pairing at the third position of the codon, allowing some tRNAs to recognize multiple codons.
Wobble Position: The 5' base of the tRNA anticodon can pair flexibly with the 3' base of the mRNA codon.
Result: Fewer tRNAs are needed to read all codons for an amino acid.
Ribosome Structure and Function
Ribosomal Subunits
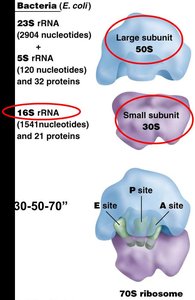
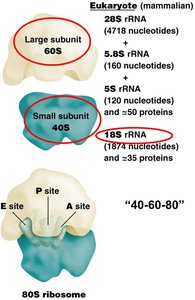
Ribosomes are composed of two subunits, each made of ribosomal RNA (rRNA) and proteins. The small subunit decodes the mRNA, while the large subunit catalyzes peptide bond formation.
Prokaryotes: 30S (small) + 50S (large) = 70S ribosome
Eukaryotes: 40S (small) + 60S (large) = 80S ribosome
Functional Sites: A (aminoacyl), P (peptidyl), and E (exit) sites for tRNA binding and movement.
Stages of Translation
Initiation
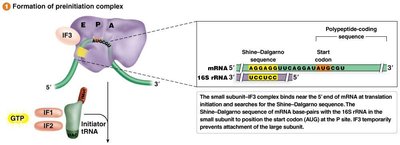
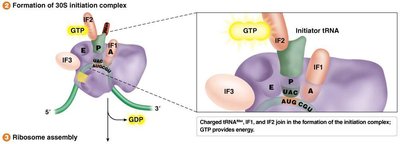
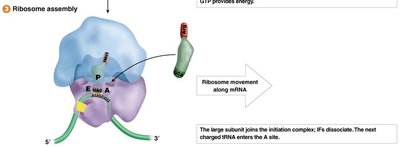
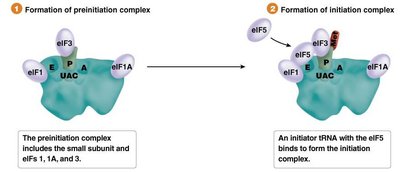
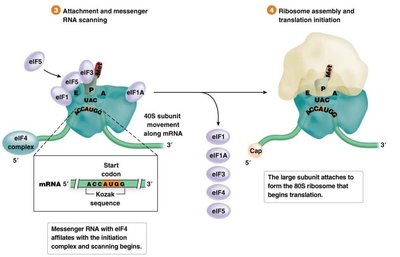
Translation initiation involves the assembly of the ribosome on the mRNA, recognition of the start codon, and recruitment of the initiator tRNA. The process differs between prokaryotes and eukaryotes.
Prokaryotes: Initiation involves the Shine-Dalgarno sequence and formylmethionine (fMet) as the initiator amino acid.
Eukaryotes: Initiation involves the 5' cap structure, scanning for the Kozak sequence, and methionine as the initiator amino acid.
Elongation
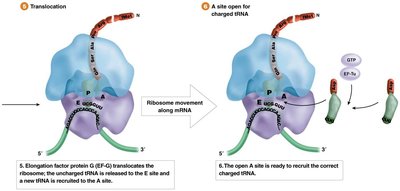
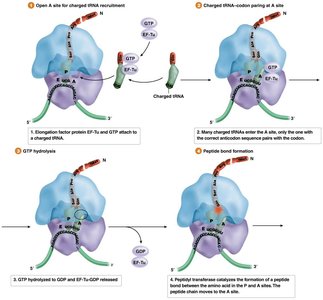
During elongation, amino acids are sequentially added to the growing polypeptide chain. This process involves the recruitment of charged tRNAs, peptide bond formation, and translocation of the ribosome along the mRNA.
Elongation Factors: Proteins that facilitate tRNA entry, peptide bond formation, and ribosome movement.
Peptidyl Transferase: An rRNA-based enzyme that catalyzes peptide bond formation.
Termination
Translation terminates when a stop codon enters the A site of the ribosome. Release factors recognize the stop codon, prompting the release of the newly synthesized polypeptide and dissociation of the ribosomal complex.
Stop Codons: UAA, UAG, UGA
Release Factors: Proteins that facilitate the release of the polypeptide chain.
Prokaryotic vs. Eukaryotic Translation
Key Differences
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Ribosome Size | 70S (30S + 50S) | 80S (40S + 60S) |
Initiation Sequence | Shine-Dalgarno | Kozak sequence |
Initiator Amino Acid | Formylmethionine (fMet) | Methionine (Met) |
Transcription/Translation Coupling | Yes | No |
Protein Structure and Function
Levels of Protein Structure
Proteins fold into specific three-dimensional structures that determine their function. The sequence of amino acids (primary structure) dictates higher levels of structure (secondary, tertiary, quaternary).
Primary Structure: Linear sequence of amino acids.
Secondary Structure: Local folding (e.g., alpha helices, beta sheets).
Tertiary Structure: Overall 3D shape of a single polypeptide.
Quaternary Structure: Association of multiple polypeptide chains.
Genetic Variation and Protein Function
Single nucleotide changes can have significant or negligible effects on protein function, depending on the location and nature of the change. Some mutations cause diseases (e.g., sickle cell anemia), while others may not affect protein function due to redundancy in the genetic code or conservative amino acid substitutions.
Additional info: This guide integrates foundational concepts from the molecular biology of translation, including the genetic code, tRNA charging, ribosome structure, and the stages of translation, with emphasis on differences between prokaryotic and eukaryotic systems. It also highlights the importance of protein structure and the impact of genetic variation on protein function.
