 Back
BackTranscription and Post-Transcriptional Processing in Eukaryotes and Prokaryotes
Study Guide - Smart Notes

Transcription: The Flow of Genetic Information
Overview of Genetic Information Flow
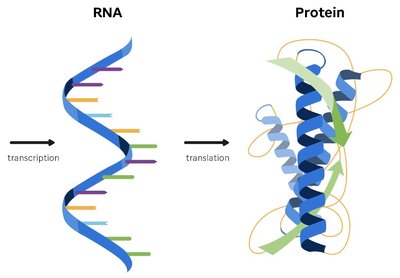
Genetic information is stored in DNA and is expressed through a two-step process: transcription and translation. Transcription is the synthesis of RNA from a DNA template, while translation is the synthesis of proteins using the information carried by mRNA.
DNA: Stores genetic information in the nucleus.
mRNA: Acts as an unstable intermediate, transferring information from the nucleus to the cytosol.
Protein Synthesis: Uses the genetic code to convert nucleotide sequences into proteins.
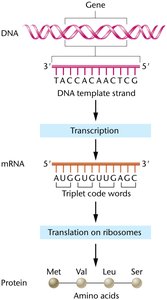
Codons: Triplet nucleotide sequences in mRNA that code for amino acids or stop signals.
Mechanism of Transcription
Chemical Reaction and Enzyme Involvement
Transcription is catalyzed by RNA polymerase, which synthesizes RNA in the 5' to 3' direction using NTPs (ATP, GTP, CTP, UTP) as substrates. Unlike DNA polymerase, RNA polymerase does not require a primer for initiation.
Template Reading: DNA template is read 3' to 5', resulting in an antiparallel RNA product.
Phosphodiester Bonds: Formed between ribonucleotides, driven by the energy from NTP cleavage.
Single-Stranded RNA: The product of transcription is a single-stranded RNA molecule.
RNA Polymerase Structure and Function
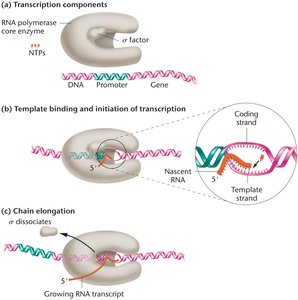
In prokaryotes, the core RNA polymerase enzyme consists of subunits α, β, β', and ω. The holoenzyme includes an additional σ (sigma) factor, which is essential for transcription initiation.
α Subunit: Scaffolding and protein interaction.
β and β': Catalytic activity and active site formation.
σ Factor: Regulatory role in promoter recognition and initiation.
Transcriptional Regulation: Promoters and Factors
Cis-Acting DNA Elements
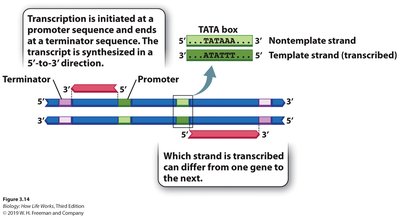
Promoters are DNA sequences located upstream (5') of the transcription start site (+1) and are essential for RNA polymerase binding and transcription initiation. Consensus sequences, such as the TATAAT (-10) and TTGACA (-35) in bacteria, are common features.
Enhancers and Silencers: Regulatory DNA elements that can be located far from the promoter, influencing transcription efficiency in eukaryotes.
Trans-Acting Factors: Proteins (e.g., RNA polymerase, transcription factors) that bind to cis-acting elements to regulate transcription.
Transcription in Prokaryotes
Initiation: σ factor recognizes the promoter, DNA unwinds, and the first ribonucleotides are linked.
Elongation: σ factor dissociates, and RNA polymerase synthesizes RNA 5' to 3'.
Termination: Specific sequences signal the end of transcription, leading to RNA release (intrinsic or Rho-dependent termination).
Transcription in Eukaryotes
Eukaryotic transcription is more complex, involving three types of RNA polymerase and extensive regulatory elements.
RNA Polymerase I: Synthesizes rRNA in the nucleolus.
RNA Polymerase II: Synthesizes mRNA and snRNA in the nucleoplasm.
RNA Polymerase III: Synthesizes 5S rRNA and tRNA in the nucleoplasm.
Form | Product | Location |
|---|---|---|
I | rRNA | Nucleolus |
II* | mRNA, snRNA | Nucleoplasm |
III | 5S rRNA, tRNA | Nucleoplasm |

TATA Box: Eukaryotic promoter element, similar to the prokaryotic -10 region.
CAAT Box: Located at -80 position, another promoter element.
Enhancers/Silencers: Can be upstream, downstream, or within introns, affecting transcription from a distance.
General Transcription Factors: Required for RNA polymerase II binding (e.g., TFIID binds TATA box).
Transcriptional Activators/Repressors: Bind enhancers/silencers to regulate gene expression.
Steps of Eukaryotic Transcription
TFIID binds to the TATA box.
Other general factors and RNA polymerase II are recruited to form the preinitiation complex.
TFIIH unwinds DNA at the promoter, allowing transcription to begin.
Elongation proceeds as general factors dissociate.
Termination occurs at a polyadenylation signal (AAUAAA), followed by mRNA cleavage.
Post-Transcriptional mRNA Processing in Eukaryotes
Summary of mRNA Processing
After transcription, eukaryotic pre-mRNA undergoes several modifications to become mature mRNA:
5′ Capping: Addition of 7-methylguanosine to the 5′ end, protecting mRNA from degradation and aiding ribosome binding.
Intron Splicing: Removal of noncoding introns and joining of exons.
Polyadenylation: Addition of a polyA tail at the 3′ end, stabilizing mRNA and facilitating export and translation.
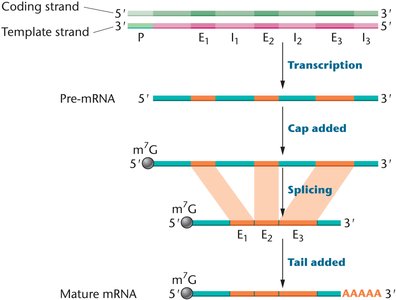
Exons and Introns
Exons: Coding sequences retained in mature mRNA.
Introns: Noncoding sequences removed during processing; allow for alternative splicing and evolutionary flexibility.
Alternative Splicing: Enables production of multiple proteins from a single gene.
Intron Removal Mechanisms
Group I Self-Splicing Introns: Catalyze their own removal via two esterification reactions; found in lower eukaryotes and plants.
Spliceosomal Introns: Removed by the spliceosome, a complex of snRNAs and snRNPs; most common in eukaryotic protein-coding genes.
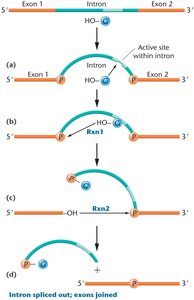
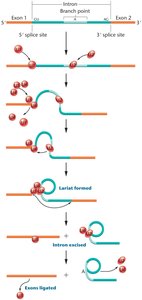
RNA Editing
Types and Functions of RNA Editing
RNA editing is a post-transcriptional process that alters the nucleotide sequence of mRNA, resulting in a different protein product than that encoded by the DNA.
Insertion/Deletion Editing: Addition or removal of nucleotides; common in plant organelles.
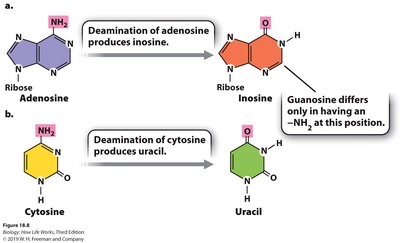
Substitution Editing: Conversion of one nucleotide to another (e.g., C to U, A to I); prevalent in animal and plant organelles.
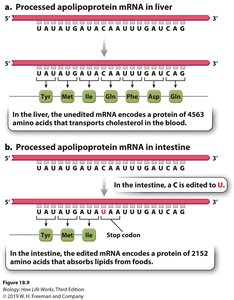
Tissue-Specific Editing: RNA editing can result in different protein products in different tissues, as seen in apolipoprotein mRNA in liver versus intestine.
Summary Table: Key Steps in Eukaryotic mRNA Processing
Step | Description | Function |
|---|---|---|
5′ Capping | Addition of 7-methylguanosine to 5′ end | Protection, ribosome binding |
Splicing | Removal of introns, joining of exons | Generates coding sequence, allows alternative splicing |
Polyadenylation | Addition of polyA tail at 3′ end | Stabilization, export, translation |
RNA Editing | Base insertion, deletion, or substitution | Protein diversity, tissue specificity |
Additional info: The notes above expand on the original lecture outline by providing definitions, mechanisms, and examples for each step of transcription and mRNA processing, as well as the regulatory elements and factors involved. The included images directly support the explanations of transcription, splicing, and RNA editing.
