 Back
BackTranscription and RNA Processing in Prokaryotes and Eukaryotes
Study Guide - Smart Notes

Transcription & RNA: Synthesis and Termination
Overview of Transcription
Transcription is the process by which RNA is synthesized from a DNA template. This process is fundamental to gene expression and is divided into three main stages: initiation, elongation, and termination. In both prokaryotes and eukaryotes, transcription is tightly regulated and involves multiple enzymes and regulatory factors.
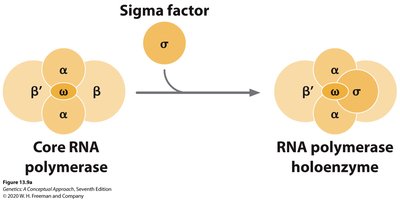
Initiation: RNA polymerase binds to the promoter region of DNA, unwinding the DNA to begin RNA synthesis.
Elongation: RNA polymerase moves along the DNA, synthesizing RNA in the 5' to 3' direction.
Termination: The process by which RNA synthesis ends and the RNA transcript is released.
Transcription Termination in Prokaryotes
Termination in prokaryotes can occur via two main mechanisms: intrinsic (Rho-independent) and Rho-dependent termination.
Intrinsic (Rho-independent) Termination
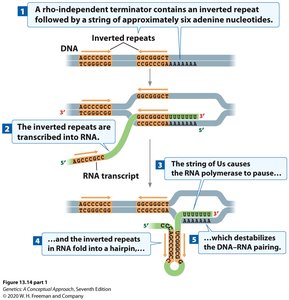
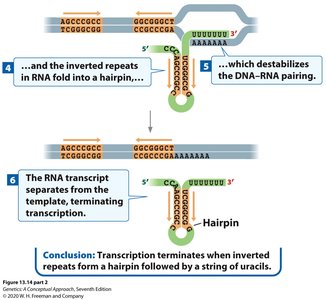
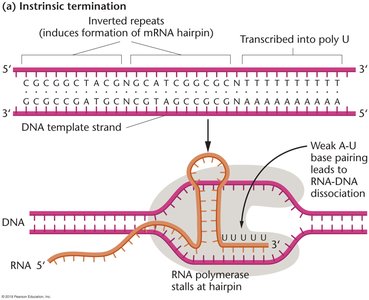
This mechanism relies on specific sequences in the DNA that, when transcribed, form a hairpin structure in the RNA followed by a string of uracils. The hairpin causes RNA polymerase to pause, and the weak A-U base pairing destabilizes the RNA-DNA hybrid, leading to transcript release.
Key features: Inverted repeats in DNA, formation of a GC-rich hairpin in RNA, followed by a poly-U tract.
Result: RNA polymerase dissociates from the DNA, releasing the RNA transcript.
Rho-dependent Termination
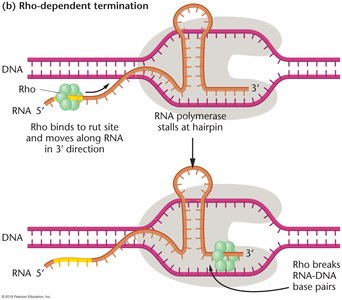
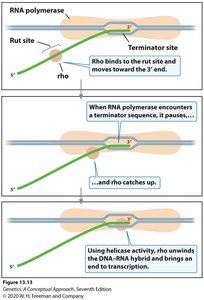
Rho-dependent termination requires the Rho (ρ) protein, which binds to a specific site on the RNA (rut site) and moves toward the RNA polymerase. When RNA polymerase stalls at a hairpin, Rho catches up and uses its helicase activity to unwind the RNA-DNA hybrid, releasing the transcript.
Key features: Rho utilization (rut) site on RNA, Rho protein, hairpin structure, no poly-U tract required.
Result: Rho protein disrupts the RNA-DNA hybrid, terminating transcription.

Transcription in Eukaryotes
RNA Polymerases in Eukaryotes
Eukaryotes possess three main RNA polymerases, each responsible for transcribing different classes of genes:
Polymerase | Product | Location |
|---|---|---|
I | rRNA | Nucleolus |
II | mRNA, snRNA | Nucleoplasm |
III | 5S rRNA, tRNA | Nucleoplasm |
Termination and Processing in Eukaryotes
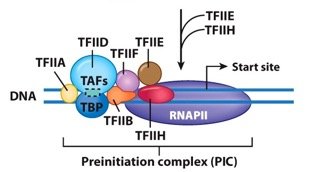
Termination mechanisms in eukaryotes differ from those in prokaryotes, especially for RNA polymerase II, where the 3' end of the transcript is defined by RNA processing rather than a specific termination sequence. Eukaryotic transcripts undergo extensive processing before becoming mature mRNA.
5' Capping: Addition of a 7-methylguanosine cap to the 5' end, which protects the RNA and aids in export and translation initiation.
3' Polyadenylation: Addition of a poly(A) tail to the 3' end, enhancing stability and export.
Splicing: Removal of non-coding introns and joining of exons to form the mature mRNA.
RNA Splicing and Processing
Splicing Mechanisms
Splicing is the process by which introns are removed from the pre-mRNA transcript. This can occur via self-splicing (Group I and II introns) or by the spliceosome (Group III introns).
Self-splicing: Certain introns can catalyze their own removal without proteins (ribozymes).
Spliceosome-dependent splicing: The spliceosome, a complex of snRNAs and proteins (snRNPs), recognizes conserved sequences at exon-intron boundaries and catalyzes intron removal.
In mammals, the 5' splice site is typically GU and the 3' splice site is AG. The average human gene contains about nine introns, which are much larger than exons.
Comparison: Prokaryotic vs. Eukaryotic Transcription
Prokaryotes: Transcription and translation are coupled; mRNA is often polycistronic and not processed.
Eukaryotes: Transcription and translation are separated by the nuclear envelope; mRNA is monocistronic and extensively processed.
Genetic Code
Codons and the Genetic Code
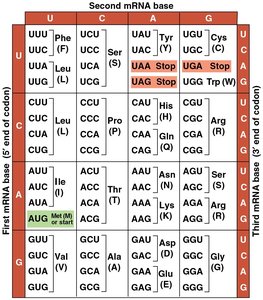
The genetic code is a set of rules by which information encoded in mRNA is translated into proteins. Each codon (a sequence of three nucleotides) specifies a particular amino acid or a stop signal for translation.
Summary Table: Key Differences in Transcription Termination
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Termination Mechanism | Intrinsic (hairpin + U's), Rho-dependent | Polymerase-specific, often via processing |
RNA Processing | Minimal | 5' capping, splicing, 3' polyadenylation |
Coupling of TC & TL | Yes | No |
Key Terms
RNA polymerase: Enzyme that synthesizes RNA from a DNA template.
Terminator: DNA sequence signaling the end of transcription.
Hairpin (stem-loop): Secondary structure in RNA important for termination.
Rho factor: Protein involved in Rho-dependent termination in prokaryotes.
Spliceosome: Complex responsible for removing introns from pre-mRNA in eukaryotes.
Exon/Intron: Coding/non-coding regions of a gene, respectively.
