 Back
BackChapter 8
Study Guide - Smart Notes

RNA Structure and Diversity
RNA Nucleotides and Structure
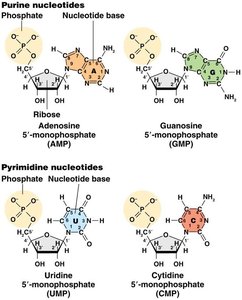
Ribonucleic acid (RNA) is a polymer composed of ribonucleotides, each consisting of a ribose sugar, a phosphate group, and one of four nitrogenous bases: adenine (A), guanine (G), cytosine (C), or uracil (U). Unlike DNA, RNA is typically single-stranded and contains uracil instead of thymine.
Purine nucleotides: Adenosine (AMP) and Guanosine (GMP)
Pyrimidine nucleotides: Uridine (UMP) and Cytidine (CMP)
Phosphodiester bonds link nucleotides in a 5' to 3' direction.
RNA Secondary Structures
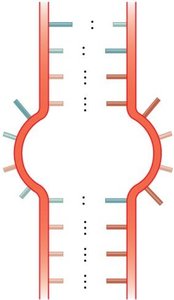
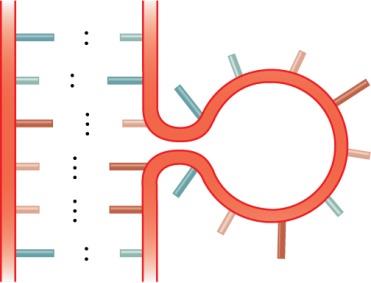
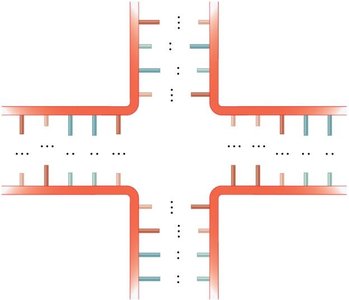
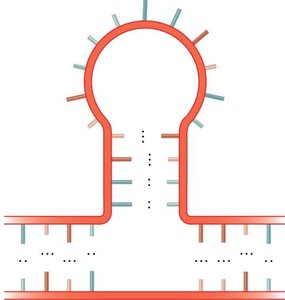
RNA molecules can fold into complex secondary structures due to intramolecular base pairing. These structures are critical for RNA function and include:
Bulge loop: Unpaired nucleotides on one side of the stem.
Internal loop: Unpaired nucleotides on both sides of the stem.
Multibranched junction: Multiple stems radiating from a single loop.
Stem-loop (hairpin): A single-stranded loop at the end of a double-stranded stem.
Transfer RNA (tRNA) Structure
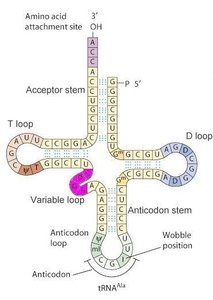
tRNA molecules have a characteristic cloverleaf secondary structure and a complex three-dimensional shape, which are essential for their role in translation.
Anticodon loop: Binds to complementary codons on mRNA.
Acceptor stem: Site for amino acid attachment.
Self-base-pairing: Maintains tRNA structure and function.
Types of RNA Molecules and Their Roles
Messenger RNA (mRNA): Encodes amino acid sequences for polypeptides.
Ribosomal RNA (rRNA): Major component of ribosomes.
Transfer RNA (tRNA): Carries amino acids to ribosomes during protein synthesis.
Small nuclear RNA (snRNA): Involved in mRNA splicing (eukaryotes).
MicroRNA (miRNA) and small interfering RNA (siRNA): Regulate gene expression (eukaryotes).
Telomerase RNA: Template for telomere elongation (eukaryotes).
Basic Mechanism of Transcription
Central Dogma of Molecular Biology
The central dogma describes the flow of genetic information: DNA is transcribed into RNA, which is then translated into protein.
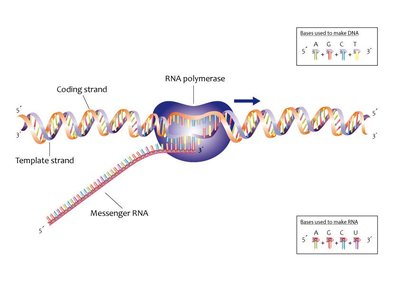
Transcription: Synthesis of RNA from a DNA template by RNA polymerase.
Template (non-coding) strand: Used by RNA polymerase to assemble a complementary RNA strand.
Coding (non-template) strand: Sequence matches the RNA (except T is replaced by U).
Bacterial Gene Transcription
Gene Structure and Regulatory Sequences
Bacterial genes are organized with specific regulatory sequences that control transcription.
Promoter: Upstream region where RNA polymerase binds to initiate transcription.
Coding region: Contains the information for protein synthesis.
Terminator: Downstream region signaling the end of transcription.
5' and 3' UTRs: Untranslated regions flanking the coding sequence in mRNA.
Ribosomal binding site (RBS): Sequence on mRNA for ribosome attachment.
Bacterial RNA Polymerase
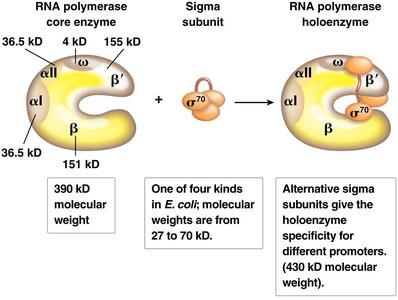
Bacterial RNA polymerase is a multi-subunit enzyme responsible for transcription.
Core enzyme: Composed of five subunits (2α, 2β, 1ω); can synthesize RNA but cannot initiate transcription alone.
Sigma (σ) subunit: Required for promoter recognition and initiation; forms the holoenzyme with the core enzyme.
Promoter Recognition and Initiation
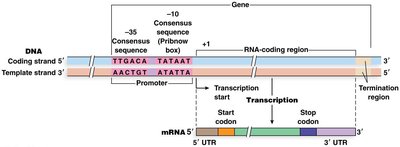
Promoters contain consensus sequences recognized by RNA polymerase and sigma factor.
-10 (Pribnow box): 5'-TATAAT-3', located ~10 bases upstream of the transcription start site.
-35 region: 5'-TTGACA-3', located ~35 bases upstream.
RNA polymerase binds these sequences, unwinds DNA, and initiates RNA synthesis.
Stages of Bacterial Transcription
1. Promoter recognition: RNA polymerase holoenzyme binds promoter.
2. Initiation: DNA unwinds, short RNA synthesized, sigma factor released.
3. Elongation: RNA polymerase synthesizes RNA 5' to 3'.
4. Termination: RNA polymerase stops and releases RNA transcript.
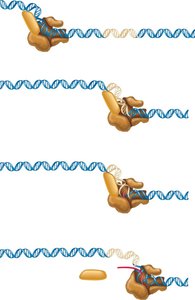
Termination Mechanisms
Transcription termination in bacteria occurs via two main mechanisms:
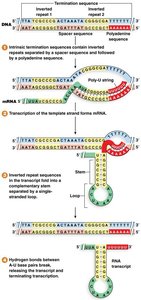
Rho-independent (intrinsic): mRNA forms a GC-rich stem-loop followed by a poly-U tail, destabilizing the RNA-DNA hybrid and causing release.
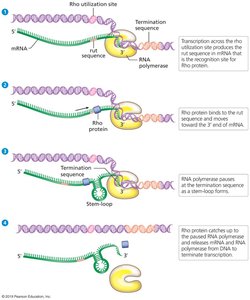
Rho-dependent: Rho protein binds to rut site on mRNA, moves toward RNA polymerase, and causes dissociation upon reaching a stem-loop structure.
Eukaryotic Gene Transcription
Complexity of Eukaryotic Transcription
Eukaryotic transcription involves more complex regulatory elements and multiple RNA polymerases.
Three RNA polymerases: Pol I (rRNA), Pol II (mRNA, snRNA, miRNA, siRNA), Pol III (tRNA, some rRNA, snRNA, miRNA, siRNA).
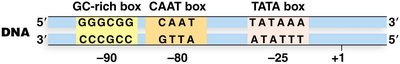
Promoters: Diverse, often containing TATA box (~-25), CAAT box (~-80), and GC-rich box (~-90).
Chromatin structure: DNA is packaged with proteins, affecting gene accessibility.
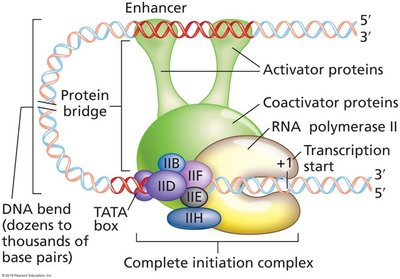
Promoter Elements and Initiation Complex
TATA box (Goldberg-Hogness box): 5'-TATAAA-3', common in Pol II promoters.
CAAT box: Near -80 position, enhances promoter activity.
GC-rich box: Near -90 or further upstream, involved in regulation.
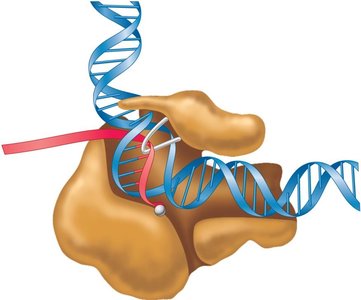
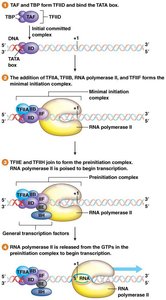
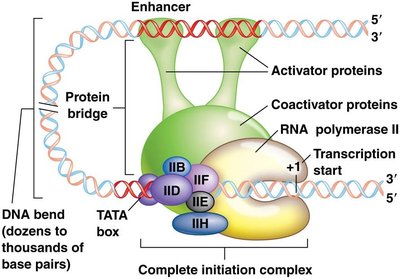
Initiation complex: General transcription factors (e.g., TFIID) and RNA polymerase assemble at the promoter.
Enhancers and Silencers
Enhancers and silencers are regulatory DNA elements that modulate transcription from a distance.
Enhancers: Bind activator proteins, increase transcription, can be upstream, downstream, or within genes.
Silencers: Bind repressor proteins, decrease transcription, also act at variable distances.
DNA bending allows interaction between distant regulatory elements and the core promoter.
Termination of Eukaryotic Transcription
Termination of transcription by RNA polymerase II involves cleavage of the RNA transcript and release of the polymerase.
Poly(A) signal sequence: Directs cleavage of the pre-mRNA.
Allosteric model: Release of elongation factors destabilizes RNA polymerase.
Torpedo model: Exonuclease degrades RNA downstream of the cleavage site, displacing RNA polymerase.
RNA Processing
Overview of RNA Processing
RNA molecules, especially in eukaryotes, undergo extensive processing before becoming functional.
5' capping: Addition of a modified guanine nucleotide to the 5' end of mRNA.
3' polyadenylation: Addition of a poly(A) tail to the 3' end of mRNA.
Splicing: Removal of introns and joining of exons in pre-mRNA.
Other modifications: tRNA and rRNA processing, RNA editing, base modification.
Eukaryotic mRNA Processing Steps
5' Capping: Protects mRNA from degradation, aids in ribosome binding, and facilitates nuclear export.
3' Poly(A) Tail: Enhances mRNA stability and translation efficiency.
Splicing: Introns are removed, and exons are ligated to form mature mRNA.
Intron Splicing Mechanism
Splicing is directed by conserved sequences at the 5' and 3' splice sites and a branch site within the intron.
5' splice site: GU sequence at the intron's 5' end.
3' splice site: AG sequence at the intron's 3' end.
Branch site: Invariant A residue upstream of the 3' splice site.
Splicing involves formation of a lariat structure and ligation of exons.
Alternative RNA Processing
Alternative processing mechanisms allow a single gene to produce multiple mRNA and protein variants.
Alternative splicing: Different patterns of exon inclusion/exclusion.
Alternative promoters: Different transcription start sites.
Alternative polyadenylation: Different poly(A) sites yield mRNAs with varied 3' ends.
This increases proteomic diversity, especially in complex organisms.
Summary Table: Key Differences in Transcription and RNA Processing
Feature | Bacteria | Eukaryotes |
|---|---|---|
RNA Polymerase | Single type, multiple sigma factors | Three types (Pol I, II, III) |
Promoter Elements | -10 (Pribnow box), -35 region | TATA box, CAAT box, GC-rich box |
Transcription/Translation | Often coupled | Physically separated (nucleus/cytoplasm) |
RNA Processing | Minimal (rare splicing, no capping/tailing) | Extensive (capping, polyadenylation, splicing) |
Introns | Rare | Common |
