 Back
BackTranscription and RNA Processing: Structure, Mechanisms, and Regulation
Study Guide - Smart Notes

RNA Structure and Diversity
RNA Structure: Chemical and Physical Properties
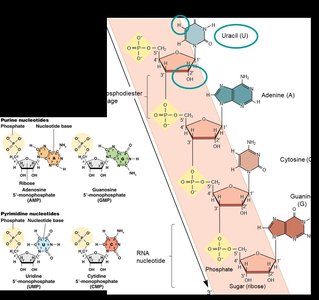
Ribonucleic acid (RNA) is a single-stranded nucleic acid that differs from DNA in several key ways. RNA contains the sugar ribose (with a hydroxyl group at the 2' carbon) and uses uracil (U) instead of thymine (T). RNA molecules can fold into complex secondary and tertiary structures, which are essential for their diverse functions in the cell.
Single-stranded nature: Allows for intramolecular base pairing, forming stems and loops.
Base pairing: Adenine (A) pairs with uracil (U), and guanine (G) pairs with cytosine (C).
Structural motifs: Common secondary structures include bulge loops, internal loops, multibranched junctions, and stem-loops.
Example: tRNA molecules exhibit extensive secondary and tertiary structure, critical for their function in translation.
RNA Secondary Structures
RNA can form a variety of secondary structures due to complementary base pairing within the same molecule. These structures are crucial for RNA stability and function.
Bulge loop: A region where one strand contains unpaired bases.
Internal loop: Unpaired bases on both strands within a double-stranded region.
Multibranched junction: A region where three or more double-stranded segments converge.
Stem-loop (hairpin): A common motif where a single-stranded loop is closed by a double-stranded stem.
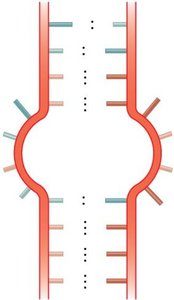
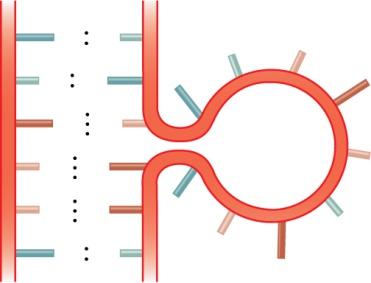
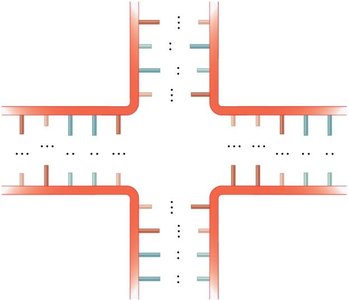
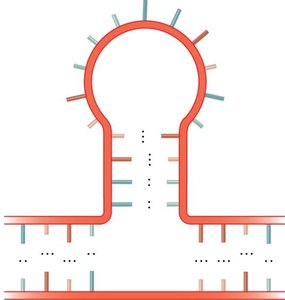
Example: The stem-loop structure is essential for the function of many non-coding RNAs and for transcription termination in bacteria.
tRNA Structure
Transfer RNA (tRNA) molecules are classic examples of RNA with complex secondary and tertiary structures. The cloverleaf secondary structure includes the acceptor stem, D loop, anticodon loop, variable loop, and T loop. The tertiary structure is L-shaped, allowing tRNA to interact with both mRNA and the ribosome during translation.
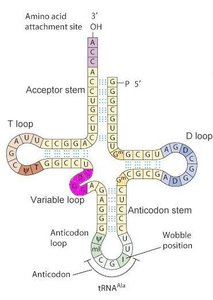
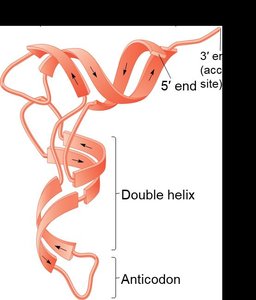
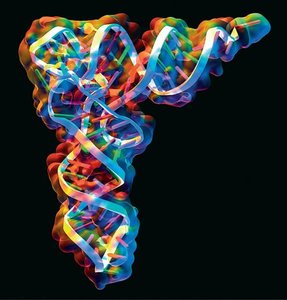
Example: The anticodon loop base-pairs with mRNA codons, while the acceptor stem attaches to specific amino acids.
Types of RNA Molecules and Their Roles
Cells contain multiple types of RNA, each with specific functions:
mRNA (messenger RNA): Encodes the sequence of amino acids in a polypeptide.
rRNA (ribosomal RNA): Major component of ribosomes, essential for protein synthesis.
tRNA (transfer RNA): Carries amino acids to ribosomes during protein synthesis.
snRNA (small nuclear RNA): Involved in splicing of pre-mRNA in eukaryotes.
miRNA (microRNA) and siRNA (small interfering RNA): Regulate gene expression by interacting with mRNA.
piRNA (piwi-interacting RNA): Protects the genome in germline cells by silencing transposable elements.
TERC (telomerase RNA): Serves as a template for telomere elongation.
Note: rRNA and tRNA are the most abundant RNA types in cells.
Basic Mechanism of Transcription
Central Dogma of Molecular Biology
The central dogma describes the flow of genetic information: DNA is transcribed into RNA, which is then translated into protein. This concept was first articulated by Francis Crick in 1958.
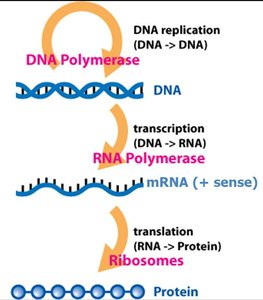
Transcription: Synthesis of RNA from a DNA template by RNA polymerase.
Translation: Synthesis of protein from an mRNA template by ribosomes.
Transcription: DNA to RNA
During transcription, RNA polymerase synthesizes a single-stranded RNA molecule using the DNA template (non-coding) strand. The coding (non-template) strand has the same sequence as the RNA (except T is replaced by U).
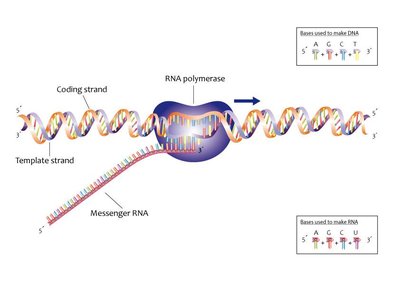
Directionality: RNA is synthesized in the 5' to 3' direction.
Base pairing: A pairs with U, G pairs with C.
Example: If the coding strand is 5'-ATGCCAATCGCATTACCG-3', the mRNA will be 5'-AUGCCAAUCGCAUUACCG-3'.
Bacterial Gene Transcription
Essential Steps of Transcription in Bacteria
Bacterial transcription involves four main steps:
Promoter recognition
Transcription initiation
Chain elongation
Chain termination
Bacterial Gene Structure
Bacterial genes are organized with regulatory sequences, a promoter, a coding region, and a terminator. The promoter is upstream of the coding region and controls RNA polymerase access.
5' and 3' UTRs: Untranslated regions flanking the coding sequence in mRNA.
Ribosomal binding site (RBS): Sequence on mRNA where ribosomes bind to initiate translation.
Bacterial RNA Polymerase
Bacterial RNA polymerase is a multi-subunit enzyme responsible for transcription. The core enzyme (α2ββ'ω) requires a sigma (σ) subunit to recognize promoters and initiate transcription.
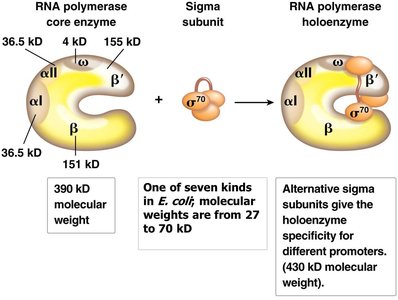
Core enzyme: Can elongate RNA but cannot initiate transcription without σ.
Holoenzyme: Core enzyme plus σ subunit, capable of promoter recognition and initiation.
Promoter Recognition and Consensus Sequences
Promoters contain conserved DNA sequences recognized by RNA polymerase. The -10 (Pribnow box, TATAAT) and -35 (TTGACA) regions are consensus sequences essential for promoter function.
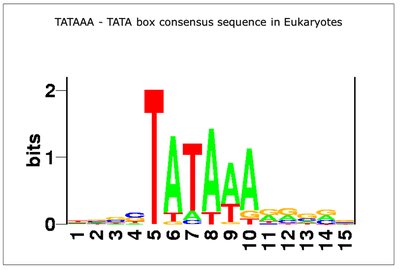
Consensus sequence: The most common nucleotide at each position in a set of similar sequences.
Transcription Initiation and Elongation in Bacteria
RNA polymerase holoenzyme binds the promoter, forming a closed complex. DNA unwinds to form an open complex, and a short RNA is synthesized. The σ factor is released, and elongation proceeds.
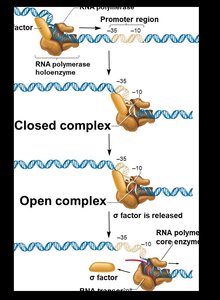
Elongation rate: ~40 nucleotides per second in E. coli.
No proofreading: Bacterial RNA polymerase lacks exonuclease activity.
Termination Mechanisms in Bacteria
Transcription termination in bacteria occurs via two main mechanisms:
Rho-independent (intrinsic): Formation of a GC-rich stem-loop followed by a poly-U tail destabilizes the RNA-DNA hybrid, causing dissociation.
Rho-dependent: The rho (ρ) protein binds a rut site on mRNA, moves toward RNA polymerase, and causes release of the transcript.
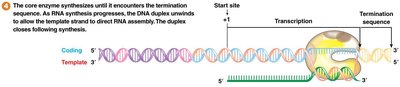
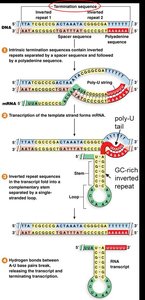
Eukaryotic Gene Transcription
Complexity of Eukaryotic Transcription
Eukaryotic transcription is more complex than bacterial transcription due to diverse promoters, regulatory proteins, the presence of introns, and chromatin structure. Eukaryotes have three RNA polymerases, each transcribing different classes of genes.
RNA Polymerase I: Transcribes rRNA genes.
RNA Polymerase II: Transcribes mRNA, snRNA, miRNA, and siRNA genes.
RNA Polymerase III: Transcribes tRNA, 5S rRNA, and some small RNAs.
Eukaryotic Promoter Elements
Promoters for RNA polymerase II are highly variable but often contain a TATA box (~-25), CAAT box (~-80), and GC-rich box (~-90). These elements are recognized by transcription factors that recruit RNA polymerase II.

TATA box: 5'-TATAAA-3', essential for accurate transcription initiation.
CAAT box: Enhances promoter strength.
GC-rich box: Involved in regulation of housekeeping genes.
Transcription Initiation in Eukaryotes
Transcription initiation involves the assembly of a large preinitiation complex at the promoter, including general transcription factors (e.g., TFIID, which contains the TATA-binding protein) and RNA polymerase II.
Enhancers: DNA elements that increase transcription levels by interacting with promoter-bound proteins, often over long distances.
Silencers: DNA elements that repress transcription by binding repressor proteins.
Eukaryotic Transcription Termination
Termination mechanisms differ among the three eukaryotic RNA polymerases. For RNA pol II, termination occurs 500-2000 nucleotides downstream of the poly-A signal. Two models explain termination: the allosteric model (polymerase destabilization) and the torpedo model (exonuclease-mediated release).
Pol I: Uses a transcription-terminating factor (TTF1).
Pol III: Terminates at a string of uracils, similar to bacterial rho-independent termination.
RNA Processing
Overview of RNA Processing
Many RNA molecules undergo processing before becoming functional. In eukaryotes, mRNA processing includes 5' capping, 3' polyadenylation, and splicing. These modifications are essential for mRNA stability, export, and translation.
5' capping: Addition of a methylated guanine (m7G) cap via a triphosphate linkage.
3' polyadenylation: Addition of a poly-A tail after cleavage downstream of the AAUAAA signal.
Splicing: Removal of introns and joining of exons by the spliceosome or self-splicing ribozymes.
Eukaryotic mRNA Processing Steps
5' Cap: Protects mRNA from degradation, aids in export, and promotes translation initiation.
3' Poly-A Tail: Increases mRNA stability and facilitates export and translation.
Splicing: Removes non-coding introns, joins coding exons.
Intron Splicing Mechanism
Splicing is catalyzed by the spliceosome, a complex of snRNPs. Introns contain conserved sequences at the 5' splice site (GU), branch site (A), and 3' splice site (AG). Splicing involves formation of a lariat structure and ligation of exons.
Self-splicing introns: Group I and II introns can catalyze their own excision (ribozymes).
Alternative RNA Processing
Alternative splicing, promoters, and polyadenylation allow a single gene to produce multiple mRNA and protein isoforms, greatly expanding proteomic diversity.
Example: The rat tropomyosin gene produces nine mRNA transcripts via alternative splicing and promoters.
Summary Table: Key Steps in Eukaryotic mRNA Processing
Step | Function |
|---|---|
5' Capping | Protects mRNA, aids export, promotes translation |
3' Polyadenylation | Stabilizes mRNA, aids export, enhances translation |
Splicing | Removes introns, joins exons |
RNA Editing | Alters nucleotide sequence post-transcriptionally |
Base Modification | Common in tRNA and rRNA for function and stability |
Additional info: In bacteria, mRNA is often translated while still being transcribed, and most bacterial mRNAs are not processed. In eukaryotes, transcription and translation are separated by the nuclear envelope, and extensive RNA processing is required before translation.
