 Back
BackTranscription: DNA to RNA – Molecular Basis and Mechanisms
Study Guide - Smart Notes

Transcription: DNA to RNA – Molecular Basis and Mechanisms
Qualities of Genetic Material
The genetic material must possess several essential qualities to ensure the faithful transmission and expression of genetic information:
Accurate Replication: The genetic material must replicate precisely so that progeny cells and organisms inherit the same genetic information as the parent.
Information Storage: It must contain stable information that governs cellular form, structure, function, development, and behavior.
Information Transfer: The material must facilitate the transfer of genetic information from DNA to RNA and ultimately to protein, as described by the Central Dogma.
The Central Dogma of Molecular Biology
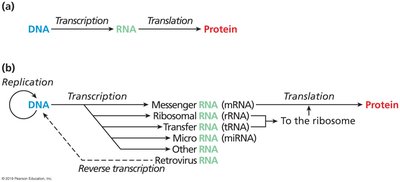
The Central Dogma, proposed by Crick (1956), describes the directional flow of genetic information:
DNA stores genetic information.
RNA acts as an intermediary, transcribing information from DNA.
Protein is synthesized from RNA, determining cellular function.
Transcription is the process of copying DNA into RNA, while translation converts RNA into protein.
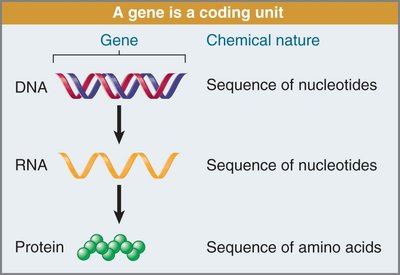
Structure and Chemical Nature of DNA and RNA
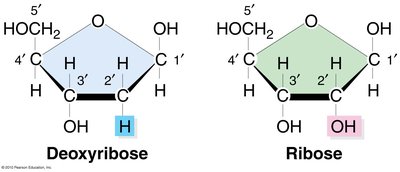
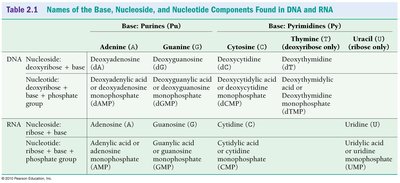
Both DNA and RNA are nucleic acids composed of nucleotide monomers, but they differ in their sugar and base composition:
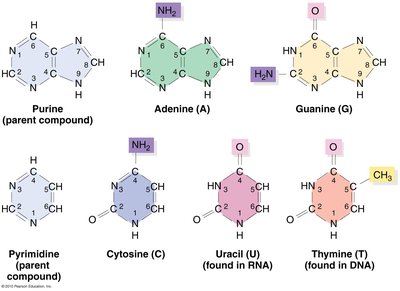
DNA: Contains deoxyribose sugar and the bases adenine (A), guanine (G), cytosine (C), and thymine (T).
RNA: Contains ribose sugar and the bases adenine (A), guanine (G), cytosine (C), and uracil (U) (instead of thymine).
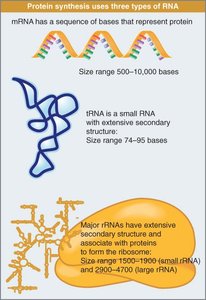
Types of RNA Involved in Protein Synthesis
Three major types of RNA participate in protein synthesis:
mRNA (Messenger RNA): Carries genetic information from DNA to the ribosome.
tRNA (Transfer RNA): Brings amino acids to the ribosome during translation.
rRNA (Ribosomal RNA): Forms the core of the ribosome's structure and catalyzes protein synthesis.
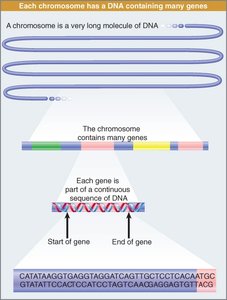
Organization of Genes and Chromosomes
Genes are located on chromosomes, which are long double-stranded DNA molecules. Each chromosome contains many genes, separated by noncoding DNA sequences:
Gene: A segment of DNA encoding a protein or functional RNA.
Chromosome: A continuous DNA molecule containing many genes.
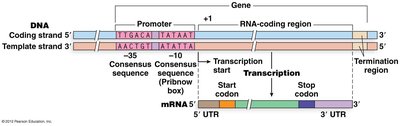
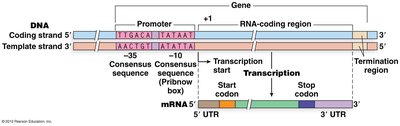
General Structure of a Gene and Transcript
A typical gene consists of regions that are transcribed and regions that are not. Transcription starts at a defined position (+1) and ends at a stop site. The resulting transcript may contain untranslated regions (UTRs) at both ends:
Promoter: Upstream region indicating where transcription begins.
Coding Region: The open reading frame (ORF) that is translated into protein.
UTRs: Sequences in mRNA that do not code for protein.
Transcription: Mechanism and Enzymes
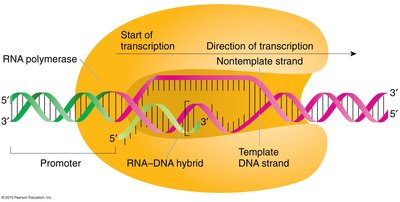
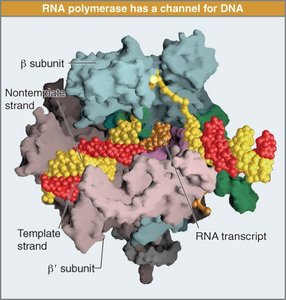
Transcription is carried out by RNA polymerases, which synthesize RNA from a DNA template:
Template Strand: One DNA strand is used to direct RNA synthesis.
Base Pairing: G pairs with C, A pairs with U in RNA (no T).
Direction: RNA synthesis occurs in the 5' to 3' direction.
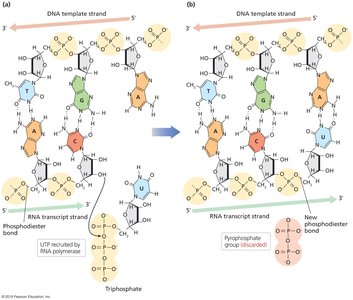
Enzyme Requirements: RNA polymerase requires NTPs (ATP, GTP, CTP, UTP) and does not need a primer.
Stages of Transcription
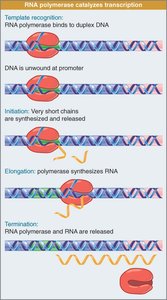
Transcription occurs in three stages:
Initiation: RNA polymerase binds to the promoter and starts RNA synthesis.
Elongation: RNA polymerase moves along DNA, extending the RNA chain.
Termination: Transcription stops and the RNA is released.
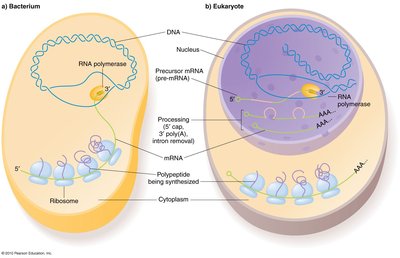
Transcription in Prokaryotes vs. Eukaryotes
There are significant differences between transcription in prokaryotes and eukaryotes:
Prokaryotes: Transcription and translation are coupled; only one RNA polymerase; no nucleus; genes are compact and uninterrupted.
Eukaryotes: Transcription occurs in the nucleus; three RNA polymerases; genes are larger, interrupted by introns; DNA is wrapped in chromatin.
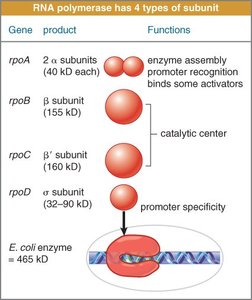
Bacterial Promoters and Sigma Factor
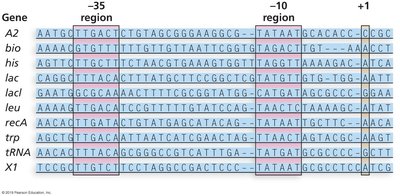
Bacterial gene promoters contain consensus sequences at the -10 and -35 positions, recognized by the sigma factor:
-10 (Pribnow box): TATAAT
-35: TTGACA
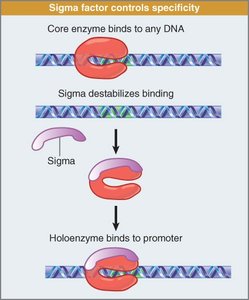
Sigma Factor: Directs RNA polymerase to the correct promoter.

Transcription Initiation and Elongation in Prokaryotes
Transcription initiation involves the formation of a closed complex, followed by local unwinding to form an open complex. Sigma factor dissociates after initiation, and core RNA polymerase continues elongation:
Closed Complex: RNA polymerase binds promoter.
Open Complex: DNA unwinds, exposing template strand.
Elongation: RNA polymerase synthesizes RNA, following base pairing rules.
Transcription Termination in Prokaryotes
Termination occurs via two mechanisms:
Intrinsic Termination: Formation of an RNA hairpin causes RNA polymerase to dissociate.
Rho-dependent Termination: Rho protein binds RNA and interacts with RNA polymerase to terminate transcription.
Transcription in Eukaryotes: Complexity and Regulation
Eukaryotic transcription is more complex, involving three RNA polymerases and multiple general transcription factors (GTFs):
RNA Polymerase I: Synthesizes rRNA.
RNA Polymerase II: Synthesizes mRNA.
RNA Polymerase III: Synthesizes tRNA and other small RNAs.
GTFs: TFIIA, TFIIB, TFIID (contains TBP), TFIIE, TFIIF, TFIIH.
Preinitiation Complex (PIC): Assembly of GTFs and RNA polymerase at the promoter.
Eukaryotic mRNA Processing
Pre-mRNAs in eukaryotes undergo three major modifications before translation:
5' Capping: Addition of a 7-methylguanosine cap to the 5' end, protecting the transcript.
Polyadenylation: Addition of a polyA tail to the 3' end, facilitating export and stability.
Splicing: Removal of introns by the spliceosome, joining exons together.
Alternative Splicing
Some eukaryotic genes undergo alternative splicing, allowing different combinations of exons to be joined, resulting in multiple protein products from a single gene.
Example: A gene with six exons may produce different proteins in different cell types by including or excluding specific exons.
Summary Table: DNA vs. RNA Components
Component | DNA | RNA |
|---|---|---|
Sugar | Deoxyribose | Ribose |
Bases | A, G, C, T | A, G, C, U |
Strands | Double-stranded | Single-stranded |
Stability | Stable | Less stable |
Key Equations
Phosphodiester Bond Formation:
Base Pairing Rules:
Additional info: These notes cover the molecular biology of transcription, including the structure and function of DNA, RNA, and proteins, the mechanisms of transcription in prokaryotes and eukaryotes, and the processing of eukaryotic mRNA. The content is directly relevant to genetics college courses, specifically chapters on the molecular biology of transcription and RNA processing.
