 Back
BackTranscription: DNA to RNA – Structure, Function, and Mechanisms
Study Guide - Smart Notes

Genetic Material: Qualities and Central Dogma
Qualities of Genetic Material
The genetic material must possess several essential qualities to ensure the faithful transmission and expression of genetic information:
Accurate Replication: The material must replicate precisely so progeny cells and organisms inherit the same genetic information as the parent.
Stable Information Storage: It must contain, in a stable form, the information that controls cellular and organismal form, structure, function, development, and behavior.
The Central Dogma of Molecular Biology
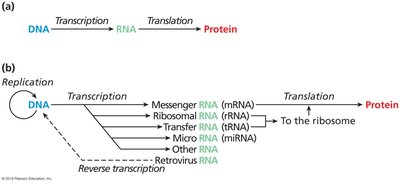
The central dogma describes the flow of genetic information within a biological system:
DNA stores genetic information.
Transcription: DNA is transcribed into RNA.
Translation: RNA is translated into protein, which determines cellular function.
Francis Crick first proposed this concept, emphasizing the directional transfer of information from DNA to RNA to protein.
Structure and Types of RNA
RNA Nucleotide Structure
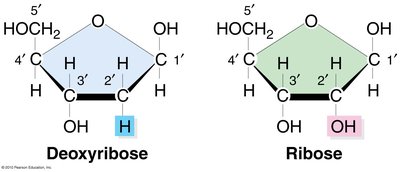
RNA is a nucleic acid composed of nucleotides, each containing:
Ribose Sugar: A five-carbon sugar with an OH group at the 2' carbon.
Phosphate Group: Attached at the 5' carbon.
Nitrogenous Base: Attached at the 1' carbon.
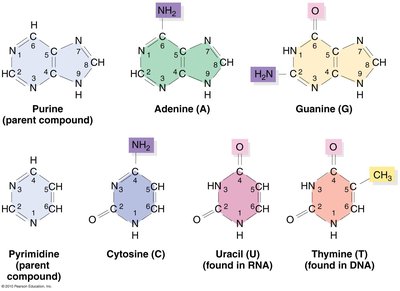
RNA Bases
RNA contains four bases: adenine (A), guanine (G), cytosine (C), and uracil (U). Uracil replaces thymine (T), which is found in DNA.
Major Types of RNA
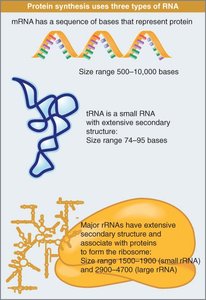
Three major types of RNA are involved in protein synthesis:
Messenger RNA (mRNA): Carries genetic information from DNA to the ribosome; highly diverse and unstable.
Transfer RNA (tRNA): Transfers amino acids to the ribosome; small and stable.
Ribosomal RNA (rRNA): Forms the core of the ribosome; abundant and stable.
Genes and Chromosomes
Organization of Genes on Chromosomes
Genes are located on chromosomes, which are long double-stranded DNA molecules. Each chromosome contains many genes separated by noncoding DNA sequences.
Genes: Encode proteins and are positioned at defined locations.
Noncoding DNA: Separates genes and may have regulatory or structural functions.
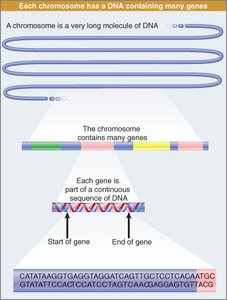
General Structure of a Gene
A gene consists of regions that are transcribed and regions that are not. Transcription starts and stops at defined positions, with the first base transcribed designated as +1.
Transcription: Mechanism and Enzymes
Transcription Process
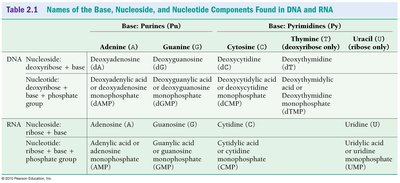
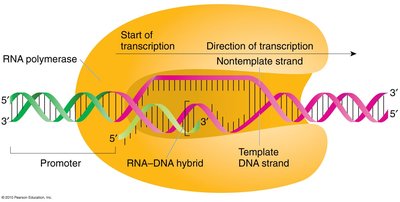
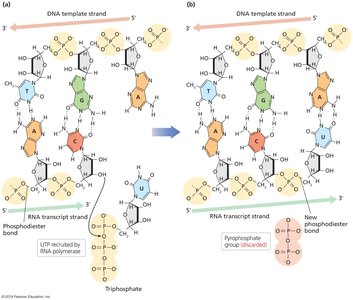
Transcription is the process by which RNA polymerase synthesizes an RNA transcript from a DNA template. Only one DNA strand (the template strand) is used, and RNA synthesis occurs in the 5' to 3' direction.
RNA Polymerase: Enzyme responsible for transcription; does not require a primer.
Base Pairing: G pairs with C, A pairs with U (in RNA).
Phosphodiester Bond Formation: RNA polymerase forms a bond between the 3' OH of the growing RNA and the 5' phosphate of the incoming nucleotide.
Stages of Transcription
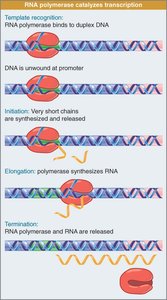
Transcription occurs in three stages:
Initiation: RNA polymerase binds to the promoter region and starts transcription.
Elongation: RNA polymerase moves along the DNA, extending the RNA chain.
Termination: Transcription stops and the RNA transcript is released.
Transcription in Prokaryotes
Bacterial RNA Polymerase and Sigma Factor
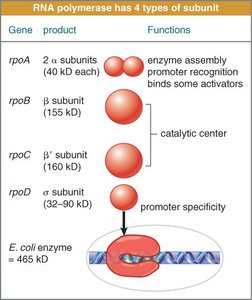
Bacterial RNA polymerase consists of multiple subunits. The sigma factor directs the core enzyme to the promoter region, forming the holoenzyme.
Core Enzyme: α2ββ′; synthesizes RNA but cannot initiate transcription alone.
Sigma Factor: Provides promoter specificity.
Promoter Regions: -10 (TATAAT) and -35 (TTGACA) consensus sequences.

Transcription Initiation and Elongation in Prokaryotes
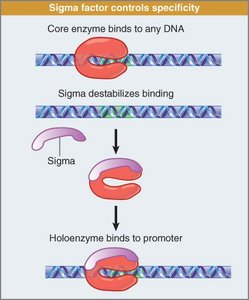
Initiation involves sigma factor binding to promoter regions, forming a closed complex. Local unwinding creates an open complex, and transcription begins. Sigma factor dissociates after a short RNA is synthesized.
Closed Complex: RNA polymerase bound to promoter, DNA double-stranded.
Open Complex: DNA unwound, template strand exposed.
Elongation: Core polymerase extends RNA transcript.
Transcription Termination in Prokaryotes
Termination occurs via intrinsic (hairpin structure) or Rho-dependent mechanisms.
Intrinsic Termination: RNA forms a hairpin, causing polymerase to dissociate.
Rho-dependent Termination: Rho protein binds RNA and interacts with polymerase to terminate transcription.
Transcription in Eukaryotes
Differences from Prokaryotes
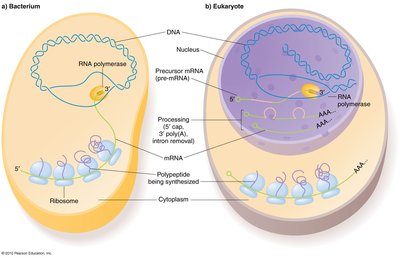
Eukaryotic transcription is more complex:
Three RNA Polymerases: I (rRNA), II (mRNA), III (tRNA and small RNAs).
Genes are larger and interrupted by introns.
Transcription occurs in the nucleus; mRNA is processed and exported to the cytoplasm.
DNA is wrapped around nucleosomes to form chromatin.
Transcription Initiation in Eukaryotes
Initiation requires general transcription factors (GTFs) that recruit RNA polymerase II to the promoter, often containing a TATA box.
TFIID: Contains TATA-binding protein (TBP); binds first to TATA box.
TFIIB, TFIIF, TFIIE, TFIIH: Sequentially recruited to form the preinitiation complex (PIC).
mRNA Processing in Eukaryotes
Eukaryotic pre-mRNAs undergo three major modifications:
5' Capping: Addition of 7-methylguanosine to the 5' end; protects from exonucleases.
Polyadenylation: Addition of a polyA tail at the 3' end; required for export and stability.
Splicing: Removal of introns by the spliceosome; exons are joined to form mature mRNA.
Alternative Splicing
Some eukaryotic genes undergo alternative splicing, allowing different combinations of exons to be joined, resulting in multiple protein products from a single gene.
Example: A gene with six exons may produce different proteins in different cell types by including or excluding specific exons.
Key Terms and Concepts
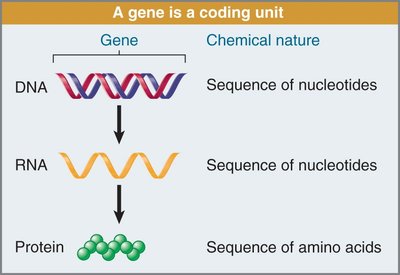
Gene: A coding unit consisting of a sequence of nucleotides in DNA.
Promoter: DNA sequence where transcription begins.
Consensus Sequence: Most common nucleotides found at each position in a collection of related DNA sites.
Introns: Noncoding sequences removed during mRNA processing.
Exons: Coding sequences retained in mature mRNA.
Table: Names of Bases, Nucleosides, and Nucleotides in DNA and RNA
Base | DNA Nucleoside | DNA Nucleotide | RNA Nucleoside | RNA Nucleotide |
|---|---|---|---|---|
Adenine (A) | Deoxyadenosine | Deoxyadenosine monophosphate (dAMP) | Adenosine | Adenosine monophosphate (AMP) |
Guanine (G) | Deoxyguanosine | Deoxyguanosine monophosphate (dGMP) | Guanosine | Guanosine monophosphate (GMP) |
Cytosine (C) | Deoxycytidine | Deoxycytidine monophosphate (dCMP) | Cytidine | Cytidine monophosphate (CMP) |
Thymine (T) | Deoxythymidine | Deoxythymidine monophosphate (dTMP) | - | - |
Uracil (U) | - | - | Uridine | Uridine monophosphate (UMP) |
Key Equations
Phosphodiester Bond Formation:
Base Pairing Rules:
Additional info: Context and examples were expanded for clarity and completeness. Images were included only when directly relevant to the explanation.
