 Back
BackTranscription in Eukaryotes: RNA Classes, Promoters, and Initiation Complexes
Study Guide - Smart Notes

Transcription in Eukaryotes
Classes of RNA in Eukaryotes
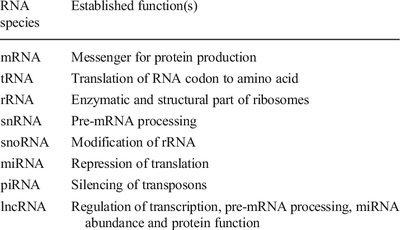
Eukaryotic cells produce a diverse array of RNA molecules, each with distinct structural and functional properties. These RNAs are classified based on their roles in gene expression and cellular function.
Messenger RNA (mRNA): Serves as the intermediary between DNA and protein synthesis; only RNA type translated into protein.
Transfer RNA (tRNA): Delivers amino acids to the ribosome during translation.
Ribosomal RNA (rRNA): Structural and enzymatic component of ribosomes.
Small Nuclear RNA (snRNA): Involved in pre-mRNA splicing within the nucleus.
Small Nucleolar RNA (snoRNA): Modifies rRNA and tRNA, found in nucleoli.
Micro RNA (miRNA): Regulates gene expression post-transcriptionally by repressing translation.
Small Interfering RNA (siRNA): Silences target mRNA by degradation.
Long Noncoding RNA (lncRNA): Regulates transcription, pre-mRNA processing, and protein function.
Piwi-interacting RNA (piRNA): Silences transposons.
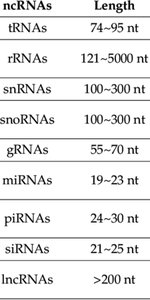
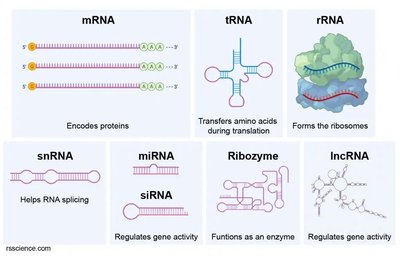
RNA Lengths: Functional RNAs vary in length, from short miRNAs (~19–23 nt) to long lncRNAs (>200 nt).

Structural Diversity: Each RNA class has a unique structure suited to its function.
Differences Between Transcription in Bacteria and Eukaryotes
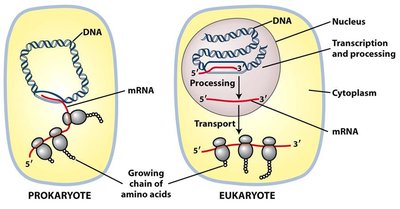
Transcription mechanisms differ significantly between prokaryotes and eukaryotes, reflecting their cellular organization and complexity.
Cellular Compartmentalization: In prokaryotes, transcription and translation occur in the same compartment; in eukaryotes, transcription occurs in the nucleus and translation in the cytoplasm.
RNA Polymerase Types: Prokaryotes have a single RNA polymerase; eukaryotes have three (Pol I, II, III).
Gene Organization: Prokaryotic genes are often in operons; eukaryotic genes are usually single transcription units.
Promoter Function: Prokaryotic promoters bind RNA polymerase directly; eukaryotic promoters bind transcription factors that regulate polymerase access.
Eukaryotic RNA Polymerases
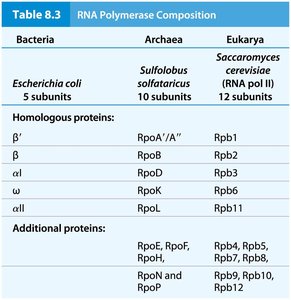
Eukaryotes utilize three distinct RNA polymerases, each responsible for transcribing specific classes of genes.
RNA Polymerase I: Transcribes most rRNA genes.
RNA Polymerase II: Transcribes protein-coding genes (mRNA) and most snRNA genes.
RNA Polymerase III: Transcribes tRNA, 5S rRNA, and some snRNA genes.
Subunit Composition: Eukaryotic RNA polymerases are structurally more complex than their bacterial counterparts, with additional subunits.
Geography of Transcription
Gene Structure and Nomenclature
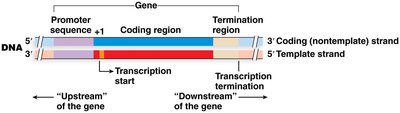
The structure of a gene includes several key regions that define the transcription process:
Promoter: Located upstream (5') of the transcription start site; controls RNA polymerase access.
Coding Region: Contains the information for the functional product (RNA or protein).
Termination Region: Located downstream (3') of the coding region; regulates transcription cessation.
+1 Nucleotide: The first nucleotide transcribed into RNA.
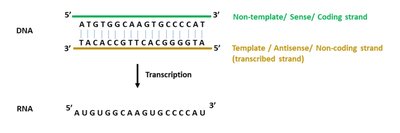
Sense and Antisense Strands
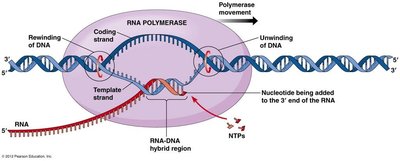
During transcription, the DNA template (antisense/non-coding) strand is read by RNA polymerase, while the coding (sense) strand has the same sequence as the resulting RNA (except for uracil replacing thymine).
Template (Antisense) Strand: Used as the template for RNA synthesis.
Coding (Sense) Strand: Matches the RNA sequence.
Promoter Elements and Transcription Factors
Promoter Sequence Elements
Promoters for RNA Polymerase II are diverse, but three elements are commonly found:
TATA Box: Consensus sequence 5'-TATAAA-3', located at -25.
CAAT Box: Often found near -80.
GC-rich Box: Consensus sequence 5'-GGGCGG-3', located at -90 or further upstream.
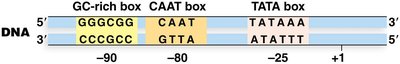
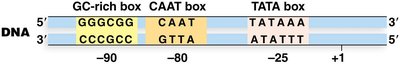
Promoters display variability in the number, type, and location of these elements.
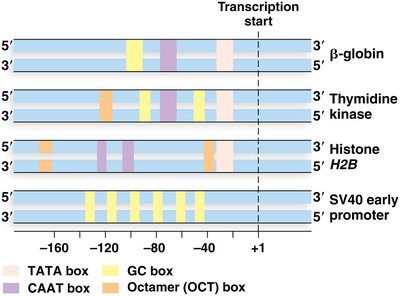
Transcription Factors
Transcription factors are proteins that bind promoter elements and regulate the accessibility of RNA polymerases:
General Transcription Factors: Required for most genes; present in all cells.
Specific Transcription Factors: Regulate expression of specific genes in a tissue- or time-specific manner.
Assembly of Transcription Initiation Complexes
Class II (RNA Polymerase II) Initiation Complex
The assembly of the transcription initiation complex at RNA Pol II promoters is a stepwise process:
Step 1: TFIID (containing TBP and TAF) binds the TATA box, forming the initial committed complex.
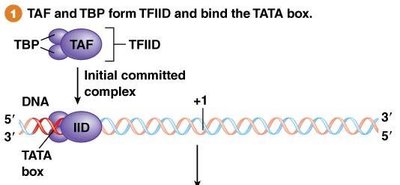
Step 2: TFIIA, TFIIB, TFIIF, and RNA Pol II join to form the minimal initiation complex.
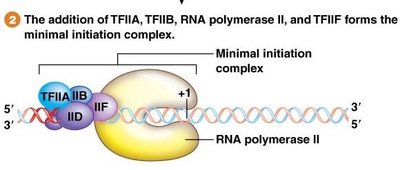
Step 3: TFIIE and TFIIH join, forming the preinitiation complex (PIC).
Step 4: RNA Pol II is directed to the +1 position to begin RNA synthesis.
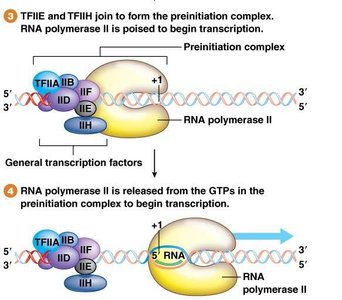
These factors are collectively known as "general transcription factors."
Enhancer and Silencer Sequences
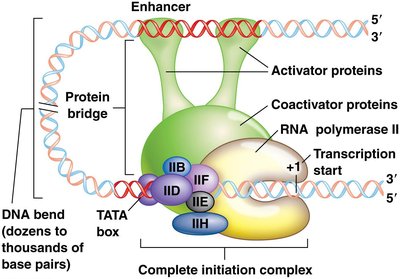
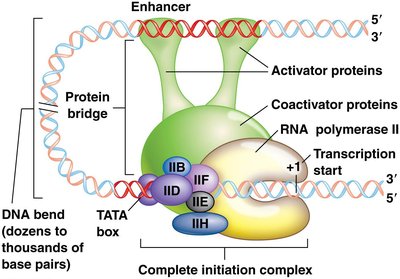
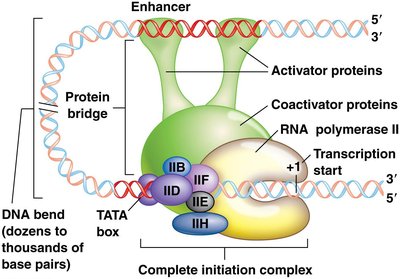
Enhancers and silencers are regulatory DNA elements that modulate gene expression at a distance:
Enhancers: Increase transcription by binding activator proteins and coactivators, forming a protein bridge to the promoter complex.
Silencers: Repress transcription by binding repressor proteins, inducing DNA bends that reduce gene expression.
Class I and Class III Transcription Initiation Complexes
Class I (RNA Polymerase I) Initiation Complex
RNA Pol I transcribes rRNA genes located in the nucleolus. Promoters recognized by Pol I have two functional elements:
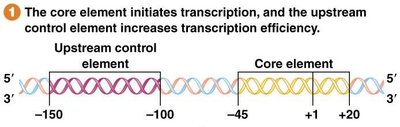
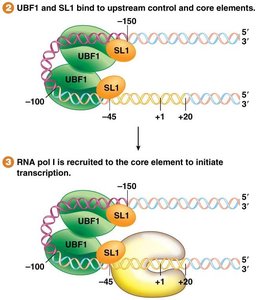
Core Element: Spans -45 to +20; required for initiation; bound by SL1 protein.
Upstream Control Element: Spans -100 to -150; increases transcription; bound by UBF1.
Class III (RNA Polymerase III) Initiation Complex
RNA Pol III transcribes tRNA, 5S rRNA, and some snRNA genes. Promoter elements differ by gene type:
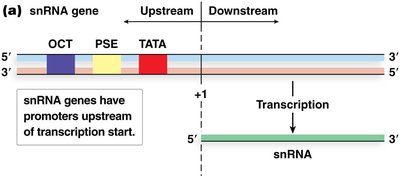
snRNA Genes: Have upstream elements (TATA box, PSE, OCT).
5S rRNA and tRNA Genes: Have internal control regions (ICRs) downstream of transcription start, called box A and box B (tRNA) or box A and box C (5S rRNA).
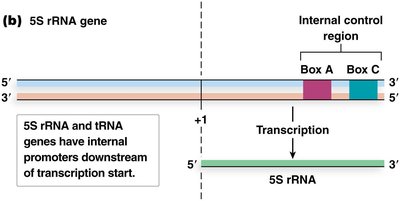
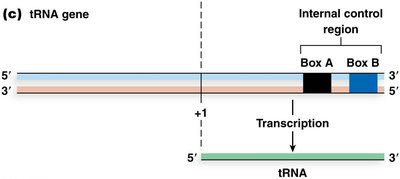
Transcription factors (TFIIIA, TFIIIC, TFIIIB) bind these elements and recruit RNA Pol III to initiate transcription.
Termination of Transcription
Termination Mechanisms
Each eukaryotic RNA polymerase utilizes a distinct mechanism for transcription termination:
RNA Pol III: Terminates at a sequence creating a string of uracils, similar to E. coli, but without a stem-loop structure.
RNA Pol I: Terminates at a 17-bp consensus sequence that binds transcription-terminating factor 1 (TTF1).
Summary Table: RNA Classes and Functions
RNA Species | Established Function(s) |
|---|---|
mRNA | Messenger for protein production |
tRNA | Translation of RNA codon to amino acid |
rRNA | Enzymatic and structural part of ribosomes |
snRNA | Pre-mRNA processing |
snoRNA | Modification of rRNA |
miRNA | Repression of translation |
piRNA | Silencing of transposons |
lncRNA | Regulation of transcription, pre-mRNA processing, miRNA abundance, and protein function |
ncRNAs | Length |
|---|---|
tRNAs | 74–95 nt |
rRNAs | 121–5000 nt |
snRNAs | 100–300 nt |
snoRNAs | 100–300 nt |
gRNAs | 55–70 nt |
miRNAs | 19–23 nt |
piRNAs | 24–30 nt |
siRNAs | 21–25 nt |
lncRNAs | >200 nt |
Additional info: These notes cover the molecular biology of transcription in eukaryotes, including RNA classes, promoter elements, transcription factors, and the assembly of initiation complexes, directly relevant to genetics college courses (Ch. 8, Ch. 13).
