 Back
BackTranscriptional and Post-Transcriptional Regulation of Gene Expression
Study Guide - Smart Notes

Transcriptional and Post-Transcriptional Regulation of Gene Expression
Overview of Gene Regulation
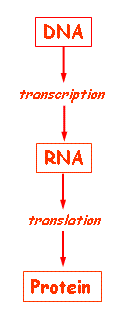
Gene expression in both prokaryotes and eukaryotes is tightly regulated at multiple levels to ensure cellular function, differentiation, and adaptation. Regulation can occur at the level of chromatin structure, transcription, mRNA processing, RNA stability, translation, and post-translational modification.
Transcriptional regulation: Determines which genes are transcribed and at what rate.
Post-transcriptional regulation: Controls mRNA processing, stability, and translation efficiency.
Translational regulation: Modulates the rate and efficiency of protein synthesis from mRNA.
Post-translational modification: Alters protein function, stability, and localization after translation.
Prokaryotic vs. Eukaryotic Genomes
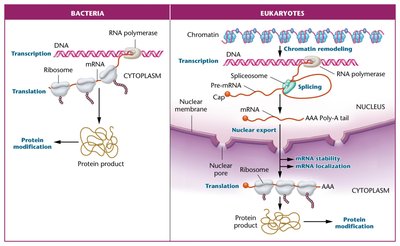
Prokaryotic and eukaryotic organisms differ significantly in genome structure and gene regulation mechanisms.
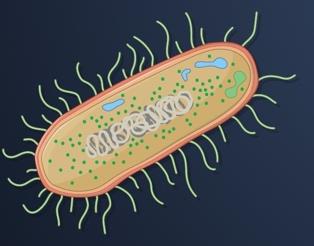
Prokaryotes (e.g., E. coli): Small, circular genomes, mostly coding DNA, no histones, rapid division and replication, minimal RNA processing.
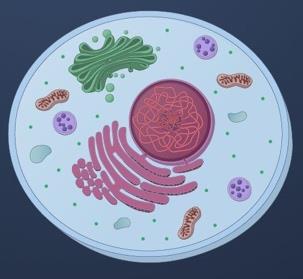
Eukaryotes (e.g., humans): Large, linear genomes, mostly noncoding DNA, DNA packaged with histones into chromatin, slower division and replication, extensive RNA processing including splicing.
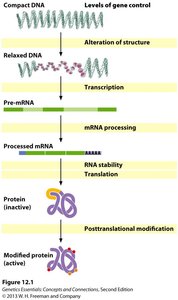
Levels of Gene Regulation
Gene regulation can occur at several hierarchical levels:
1. DNA or chromatin structure alteration
2. Transcription regulation
3. Post-transcriptional regulation (mRNA processing and stability)
4. Translation
5. Post-translational modification
1. DNA or Chromatin Structure Alteration
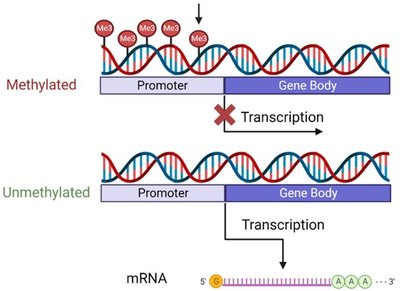
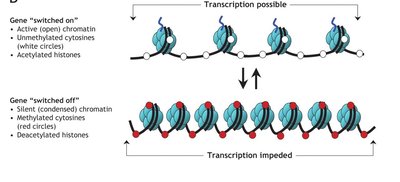
DNA Methylation
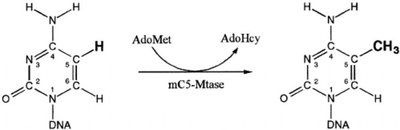
DNA methylation involves the addition of a methyl group to cytosine bases, typically at CpG islands, leading to transcriptional repression. This modification is tissue-specific and contributes to chromatin silencing.
CpG islands: Regions rich in cytosine and guanine nucleotides, often found near gene promoters.
DNA methyltransferases: Enzymes that catalyze the transfer of methyl groups to DNA.
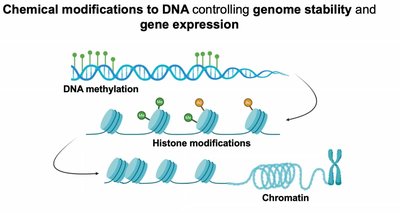
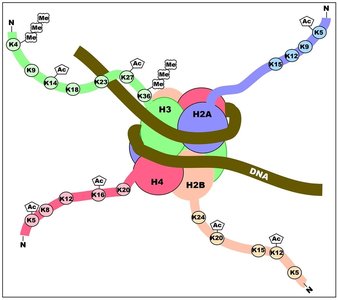
Histone Modification
Histone proteins can be modified by methylation, acetylation, phosphorylation, and ubiquitination, affecting chromatin structure and gene accessibility.
Histone methylation: Can activate or repress transcription depending on the site and number of methyl groups.
Histone acetylation: Generally associated with transcriptional activation by loosening chromatin structure.
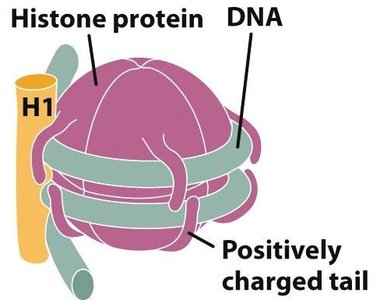
Chromatin Remodeling
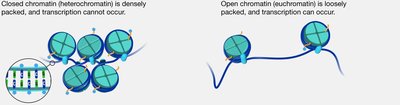
Chromatin-remodeling complexes reposition nucleosomes, making DNA more or less accessible to transcription machinery. Chromatin can exist as:
Heterochromatin: Densely packed, transcriptionally inactive.
Euchromatin: Loosely packed, transcriptionally active.
2. Transcription Regulation
Gene Regulatory Architecture
Transcription regulation involves the interaction of DNA elements and proteins to control gene expression. Key components include:
Core promoter: Region immediately upstream of the gene, necessary for transcription initiation.
Basal transcription apparatus: Includes RNA polymerase and general transcription factors.
Distal regulatory elements: Enhancers, silencers, and insulators that can be located far from the gene.
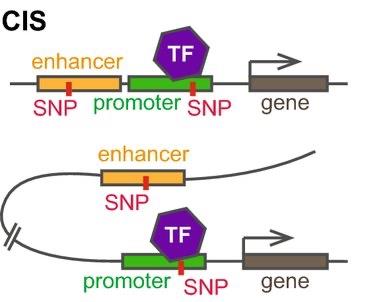
Cis- and Trans-Regulatory Elements
Cis-acting regulatory elements (CREs): Non-coding DNA sequences that regulate neighboring genes (e.g., promoters, enhancers, silencers, insulators).
Trans-regulatory elements (TREs): Proteins or RNAs that regulate gene expression from a distance (e.g., transcription factors, RNA polymerase II).

Promoters
Promoters are DNA regions upstream of the transcription start site (TSS) that recruit RNA polymerase and transcription factors. They contain consensus sequences such as the TATA box and CAAT box.
Each gene may have multiple promoters and TSSs.
Promoters determine the binding of RNA polymerase and the initiation of transcription.
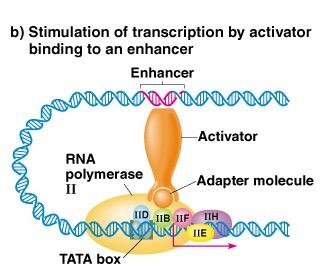
Enhancers
Enhancers are DNA sequences that stimulate transcription from a distance. They function by looping the DNA to bring activator-bound enhancers close to promoters, facilitating transcription initiation.
Can be located upstream, downstream, or within introns.
Orientation-independent and can act over long distances.
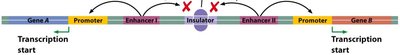
Silencers and Insulators
Silencers are DNA elements that repress transcription, while insulators block the interaction between enhancers and promoters, creating boundaries between regulatory domains.
Silencers: Bind repressor proteins to inhibit transcription.
Insulators: Bind proteins like CTCF to block enhancer activity and protect genes from heterochromatin spread.

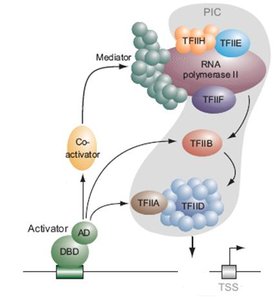
Transcription Factors (TFs)
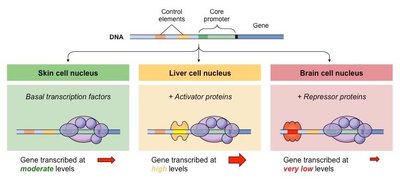
Transcription factors are proteins that bind specific DNA sequences to regulate transcription. They can act as activators or repressors and are often context-specific.
Activator proteins increase transcription by stabilizing the transcription machinery.
Repressor proteins decrease transcription by blocking activator binding or basal machinery assembly.

3. Post-Transcriptional Regulation
mRNA Processing
In eukaryotes, pre-mRNA undergoes several processing steps before becoming mature mRNA:
5' capping: Addition of a methylated guanine cap to the 5' end.
3' polyadenylation: Addition of a poly(A) tail to the 3' end.
Splicing: Removal of introns and joining of exons; alternative splicing allows for multiple protein isoforms from a single gene.
RNA Stability and Degradation
The steady-state level of mRNA is determined by the balance between transcription and degradation. Mechanisms include deadenylation, decapping, and exonucleolytic decay.
mRNA stability varies widely, influencing gene expression levels.
RNA Interference (RNAi)
RNAi is a post-transcriptional gene silencing mechanism mediated by small non-coding RNAs (siRNAs and miRNAs). These RNAs guide the RNA-induced silencing complex (RISC) to complementary mRNAs, leading to their degradation or translational repression.
Dicer: Enzyme that processes double-stranded RNA into siRNA or miRNA.
RISC: Protein complex that mediates mRNA silencing.
4. Translational Regulation
Translational regulation affects the rate at which proteins are synthesized from mRNA. Factors include mRNA localization, availability of ribosomes and initiation factors, and the action of miRNAs that can block translation initiation.
miRNAs can bind to the 5' cap or 3' UTR of mRNAs, preventing translation initiation.
5. Post-Translational Modification (PTM)
PTMs alter protein function, stability, and localization after translation. Common modifications include acetylation, phosphorylation, methylation, ubiquitination, and glycosylation.
Ubiquitin-mediated degradation: Proteins tagged with ubiquitin are targeted for degradation by the proteasome.
Summary Table: Types and Actions of Gene Regulation
Regulation Type | Action Type | Level of Regulation |
|---|---|---|
Chromatin Remodeling | Activate | Chromatin Structure |
Histone Methylation | Activate/Inhibit | Chromatin Structure |
Histone Acetylation | Activate | Chromatin Structure |
DNA Methylation | Inhibit | DNA Structure |
Activators | Activate | Transcription |
Repressors | Inhibit | Transcription |
Enhancers | Activate | Transcription |
Insulators | Inhibit | Transcription |
RNA Splicing | Activate/Inhibit | mRNA Processing |
RNA Degradation | Inhibit | RNA Stability |
Interfering RNA (siRNA, miRNA) | Inhibit | RNA Stability |
Translational Apparatus Availability | Activate/Inhibit | Translation |
Post-translational Modifications | Activate/Inhibit | PTM |
Key Concepts for Exam Preparation
Understand the different levels of gene regulation and their molecular mechanisms.
Be able to compare prokaryotic and eukaryotic gene regulation.
Know the roles of cis- and trans-regulatory elements, transcription factors, and chromatin modifications.
Be familiar with mRNA processing, RNA interference, and post-translational modifications.
