 Back
BackTranslation and Post-Translational Modifications in Genetics
Study Guide - Smart Notes

Translation: Decoding the Genetic Message
Overview of Translation
Translation is the process by which the genetic information encoded in messenger RNA (mRNA) is used to synthesize polypeptides (proteins). This process occurs in the cytoplasm and involves ribosomes, transfer RNAs (tRNAs), and various protein factors. The genetic code, composed of nucleotide triplets called codons, specifies the sequence of amino acids in a protein.
Key Components: mRNA, tRNA, ribosomes, amino acids, and translation factors.
Stages of Translation: Initiation, elongation, and termination.
Genetic Code: The code is degenerate, meaning multiple codons can specify the same amino acid.
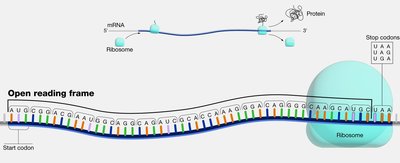
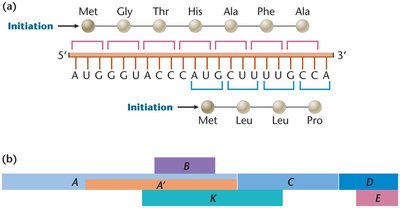
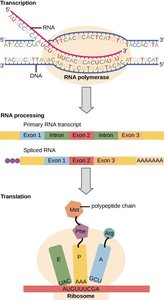
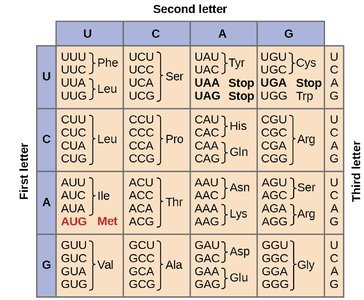
Open Reading Frame (ORF)
An open reading frame (ORF) is a continuous stretch of nucleotides in mRNA that begins with a start codon (usually AUG) and ends with a stop codon (UAA, UAG, or UGA). ORFs are the regions that can be translated into proteins.
Start Codon: Typically AUG, coding for methionine.
Stop Codons: UAA, UAG, UGA; signal the end of translation.
Translation Machinery
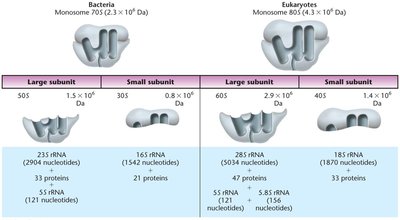
Translation requires the coordinated action of ribosomes, tRNAs, and various protein factors. Ribosomes are large complexes of rRNA and proteins, and they provide the site for mRNA decoding and peptide bond formation.
Prokaryotic Ribosomes: 70S (50S large subunit + 30S small subunit)
Eukaryotic Ribosomes: 80S (60S large subunit + 40S small subunit)
tRNA Structure and Function
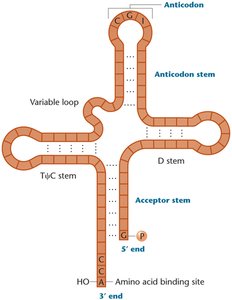
Transfer RNAs (tRNAs) are small, stable RNA molecules that serve as adapters, matching amino acids to their corresponding codons in mRNA. Each tRNA has an anticodon loop that base-pairs with the mRNA codon and an acceptor stem where the amino acid is attached.
Cloverleaf Structure: Four stems and three loops, with the anticodon loop recognizing the mRNA codon.
CCA Tail: The 3' end of tRNA always ends with CCA, the attachment site for amino acids.
Charging of tRNAs
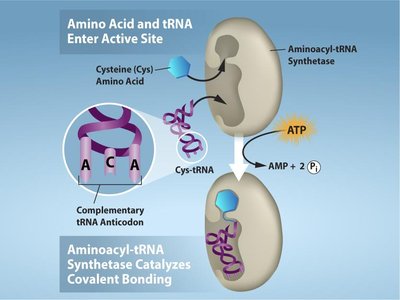
Aminoacyl-tRNA synthetases are enzymes that attach the correct amino acid to its corresponding tRNA, a process called "charging." There are 20 different synthetases, one for each amino acid, and this process requires ATP.
Specificity: Each synthetase recognizes only one amino acid and its compatible tRNAs.
Energy Requirement: ATP is hydrolyzed to AMP and PPi during the charging reaction.
tRNA Modifications
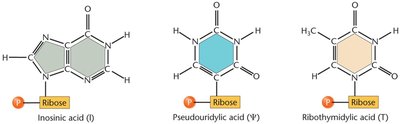
tRNAs undergo extensive post-transcriptional modifications, including base modifications, trimming of precursor sequences, addition of the CCA tail, and removal of introns. These modifications enhance tRNA stability and accuracy during translation.
Modified Bases: Inosine, pseudouridine, ribothymidine, etc.
Function: Improve stability, increase translation accuracy, and allow flexibility in codon recognition.
Ribosomal DNA (rDNA) and rRNA Genes
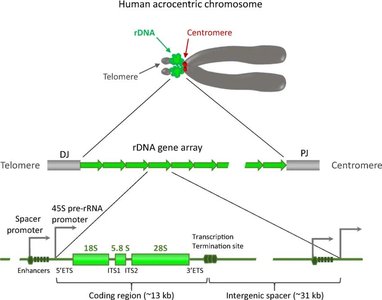
Ribosomal DNA (rDNA) encodes ribosomal RNA (rRNA) molecules, which are essential components of ribosomes. rDNA genes are organized in tandem repeats and are found in clusters at specific chromosomal regions called nucleolus organizer regions (NORs).
rRNA Genes: Encode 18S, 5.8S, and 28S rRNAs in eukaryotes.
Spacer DNA: Noncoding sequences separate rRNA genes and help regulate transcription.
Stages of Translation
Initiation (Prokaryotes)
The small 30S ribosomal subunit binds to the mRNA at the Shine–Dalgarno sequence, aligning the start codon (AUG).
The initiator tRNA carrying N-formylmethionine (fMet) binds to the P site.
Initiation factors (IF1, IF2, IF3) assist in complex formation.
The large 50S subunit joins, and GTP is hydrolyzed to release initiation factors.
Elongation (Bacteria)
Charged tRNAs enter the A site, peptide bonds form, and the ribosome translocates along the mRNA.
Elongation factors (EF-Tu, EF-G) facilitate tRNA entry and ribosome movement.
Polysomes: Multiple ribosomes can translate a single mRNA simultaneously.
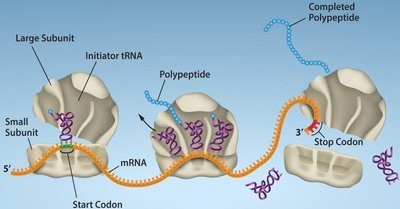
Termination
Stop codons (UAA, UAG, UGA) in the A site signal termination.
Release factors (RF1, RF2, RF3) recognize stop codons and release the polypeptide.
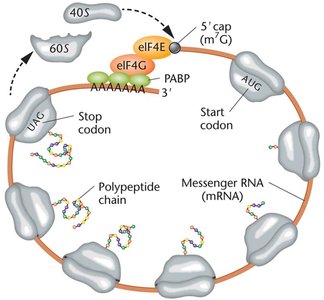
Translation in Eukaryotes
Ribosomes are larger (80S) and translation is more complex.
mRNAs are capped at the 5' end and polyadenylated at the 3' end.
The Kozak sequence surrounds the start codon, enhancing initiation efficiency.
Initiation factors (eIFs) and poly-A binding proteins (PABPs) are involved.
One-Gene–One-Enzyme and One-Gene–One-Polypeptide Hypotheses
Historical Perspective
The one-gene–one-enzyme hypothesis, proposed by Beadle and Tatum, established that each gene encodes a specific enzyme. This was later refined to the one-gene–one-polypeptide hypothesis, recognizing that not all proteins are enzymes and that some proteins are composed of multiple polypeptide chains.
Example: Mutations in the gene for tyrosine biosynthesis in Neurospora block growth on minimal medium unless tyrosine is supplied.
Protein Folding, Modification, and Targeting
Protein Folding
After translation, polypeptides fold into specific three-dimensional structures. Some proteins fold spontaneously, while others require chaperone proteins to assist in proper folding. Incorrect folding can lead to dysfunctional proteins and disease.
Signal Sequences: Short amino-terminal sequences direct proteins to specific cellular compartments.
Chaperones: Assist in folding and prevent aggregation.
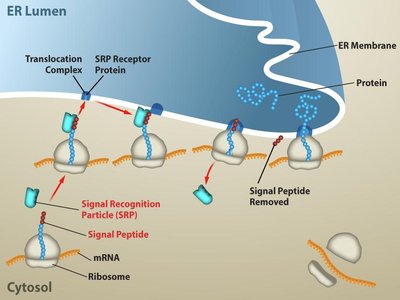
Protein Targeting
Proteins destined for secretion or specific organelles contain signal sequences that are recognized by signal recognition particles (SRPs), directing them to the endoplasmic reticulum (ER) or other locations.
Peptide Bond Formation and Protein Structure
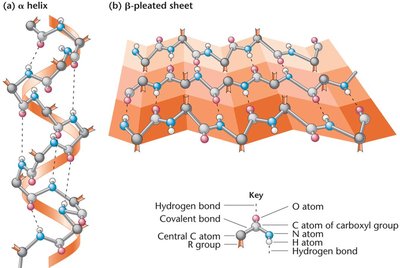
Peptide bonds are formed by dehydration reactions between the carboxyl group of one amino acid and the amino group of another. Proteins have four levels of structure: primary (sequence), secondary (α-helix, β-sheet), tertiary (3D conformation), and quaternary (multiple polypeptide chains).
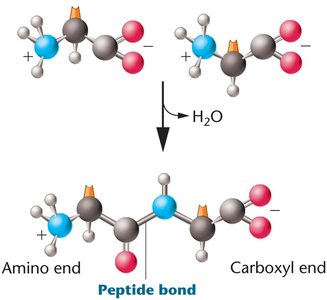
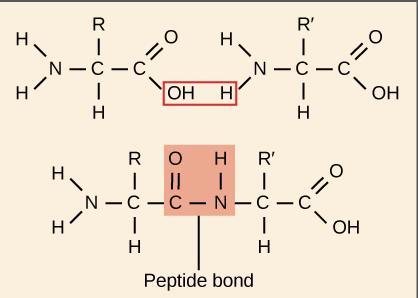
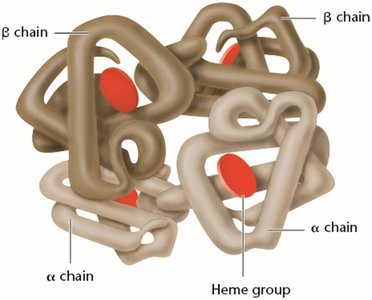
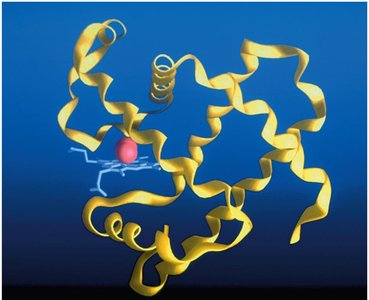
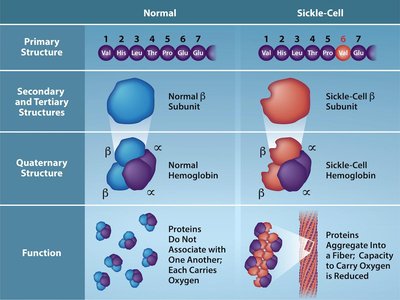
Genetic Mutations and Protein Function
Mutations in the coding sequence of a gene can alter the amino acid sequence of the resulting protein, potentially affecting its structure and function. For example, a single nucleotide change in the β-globin gene causes sickle cell anemia by substituting valine for glutamic acid.
Point Mutation: Single nucleotide change (e.g., GAG → GTG in sickle cell disease).
Frameshift Mutation: Insertions or deletions that alter the reading frame, often resulting in nonfunctional proteins.
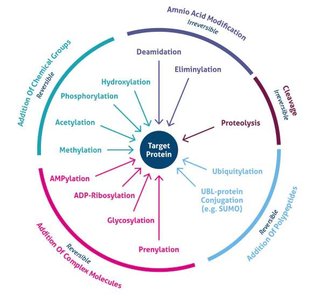
Post-Translational Modifications
Types and Functions
After translation, proteins often undergo post-translational modifications (PTMs) that are essential for their function, stability, localization, and regulation. Common PTMs include phosphorylation, acetylation, methylation, glycosylation, ubiquitination, SUMOylation, lipidation, hydroxylation, and disulfide bond formation.
Modification | Main Function |
|---|---|
Phosphorylation | Regulates enzyme activity and cell signaling |
Acetylation | Controls gene expression and protein stability |
Methylation | Modulates protein interactions and gene regulation |
Glycosylation | Aids protein folding, stability, and cell recognition |
Ubiquitination | Tags proteins for degradation by the proteasome |
SUMOylation | Regulates protein localization and transcription |
Lipidation | Targets proteins to cell membranes |
Hydroxylation | Important for collagen stability and oxygen sensing |
Disulfide bonds | Stabilize protein structure |
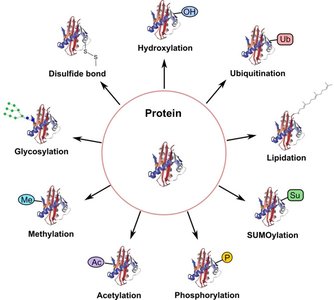
Protein Domains and Functional Organization
Protein Domains
Protein domains are distinct structural and functional units within a protein, typically consisting of 50–300 amino acids. Domains can confer specific functions, such as DNA binding, dimerization, or ligand binding, and allow proteins to participate in diverse cellular processes.
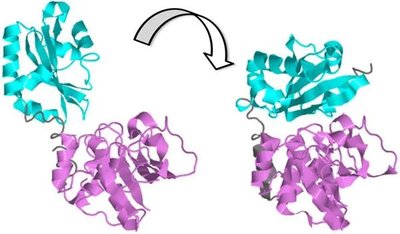
Protein-Protein Interactions (PPI)
Protein-protein interactions are essential for many cellular functions, including signal transduction, gene regulation, and metabolic pathways. These interactions often induce conformational changes that activate or inhibit protein function.
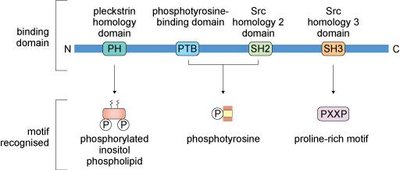
Signaling Domains
Signaling proteins contain conserved domains that recognize specific motifs, such as phosphorylated tyrosine or proline-rich sequences. These domains facilitate interactions with other proteins, lipids, or regulatory factors, enabling complex signaling networks.
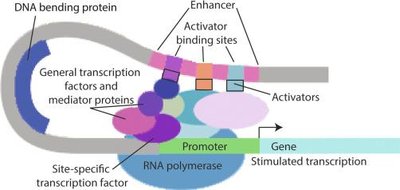
Regulation of Gene Expression
Overview
Gene expression is regulated at multiple levels, from chromatin structure and transcription to mRNA processing, translation, and protein degradation. The final protein level in the cell depends on both synthesis and degradation rates.
Ubiquitination and Protein Degradation
Ubiquitination is a post-translational modification that tags proteins for degradation by the proteasome. This process is crucial for regulating protein levels and controlling gene expression.
Polyubiquitination (K48-linked): Classical signal for proteasomal degradation.
Other Ubiquitin Linkages: Can regulate signaling or localization without degradation.
Example: NF-κB Pathway Regulation
The NF-κB signaling pathway is regulated by multiple post-translational mechanisms, including ubiquitination, phosphorylation, proteolytic cleavage, and translocation of transcription factors into the nucleus. These modifications control the activation and inactivation of inflammatory gene expression.
Summary Table: Key Steps and Components in Translation
Step | Key Components | Function |
|---|---|---|
Initiation | Small ribosomal subunit, initiator tRNA, initiation factors | Assembly of translation complex at start codon |
Elongation | Charged tRNAs, elongation factors, ribosome | Sequential addition of amino acids to polypeptide chain |
Termination | Stop codon, release factors | Release of completed polypeptide |
Post-Translational Modification | Enzymes (kinases, transferases, etc.) | Modification of protein structure and function |
Additional info: These notes integrate foundational concepts from translation, protein structure, and gene regulation, providing a comprehensive overview suitable for exam preparation in a college genetics course.
