 Back
BackTranslation and Protein Structure: From Genes to Functional Proteins
Study Guide - Smart Notes

Translation and Proteins
Overview of Translation
Translation is the process by which the genetic code carried by mRNA is decoded to produce a specific sequence of amino acids, resulting in the synthesis of a polypeptide chain. This process is fundamental to gene expression and is tightly regulated in both prokaryotic and eukaryotic cells.
Polyribosomes (Polysomes)
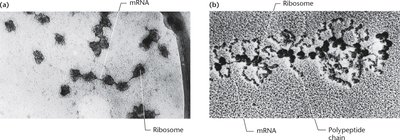
Polyribosomes are complexes of multiple ribosomes simultaneously translating a single mRNA molecule. This arrangement increases the efficiency of protein synthesis by allowing multiple copies of a protein to be produced from a single mRNA transcript.
Structure: Multiple ribosomes attached to a single mRNA strand.
Function: Enables rapid and efficient translation of proteins.
Observation: Polypeptide chains can be seen emerging from ribosomes in electron micrographs.
Prokaryotic vs. Eukaryotic Translation
Key Differences
Translation differs significantly between prokaryotes and eukaryotes in terms of ribosome structure, mRNA processing, and regulatory mechanisms.
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Ribosome Size | Smaller (70S) | Larger (80S) |
mRNA Half-life | Minutes | Hours |
Spatial/Temporal Control | Transcription and translation are coupled | Transcription in nucleus, translation in cytoplasm |
Initiation Sequence | Shine-Dalgarno sequence | Kozak sequence, 5' m7G cap |
Initiating Amino Acid | Formylmethionine (fMet) | Methionine (Met) via tRNAiMet |
mRNA Structure | Polycistronic common | Monocistronic common |
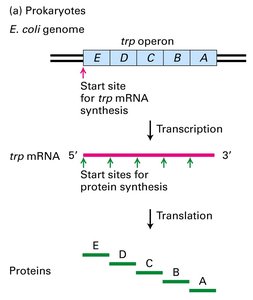
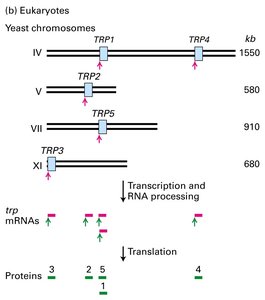
Closed-Loop Translation in Eukaryotes
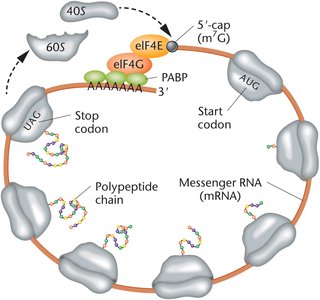
In eukaryotes, the 3′ polyA tail and the 5′ m7G cap of mRNA interact via protein factors to form a closed-loop structure. This configuration enhances translation efficiency and protects mRNA from degradation.
PolyA binding proteins (PABP): Bind the polyA tail and interact with initiation factors.
Initiation factors: Bind the 5′ cap and facilitate ribosome recruitment.
Advantages: Prevents translation of degraded mRNA and promotes ribosome recycling.
Protein Synthesis Inhibitors
Antibiotics and Inhibitors
Several antibiotics and chemical inhibitors target different steps of protein synthesis, providing tools for research and clinical therapy.
Prokaryote-specific: Tetracycline, Streptomycin, Chloramphenicol, Erythromycin, Rifamycin
Both prokaryotes and eukaryotes: Puromycin, Actinomycin D
Eukaryote-specific: Cycloheximide, Anisomycin
These inhibitors act by blocking tRNA binding, ribosome movement, peptide bond formation, or RNA synthesis.
One Gene: One Enzyme and Beyond
Historical Evidence: Alkaptonuria
Alkaptonuria is an inherited metabolic disorder where homogentisic acid is not metabolized due to a defective enzyme, leading to its excretion and blackening of urine. This provided early evidence that genes encode enzymes.
Beadle and Tatum's Experiments
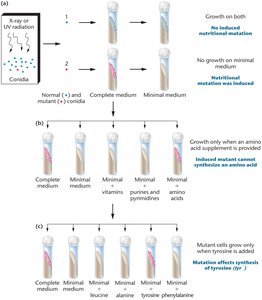
Beadle and Tatum used the bread mold Neurospora to demonstrate that mutations in specific genes disrupt individual steps in metabolic pathways, supporting the "one gene: one enzyme" hypothesis.
Mutants unable to grow on minimal medium were rescued by supplementing with specific nutrients, indicating a block in a single enzymatic step.
One Gene: One Polypeptide Chain
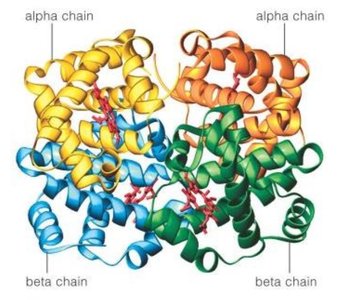
Further research revealed that not all proteins are enzymes and many proteins are composed of multiple polypeptide chains. The modern view is "one gene: one polypeptide chain." For example, hemoglobin consists of multiple subunits, each encoded by a separate gene.
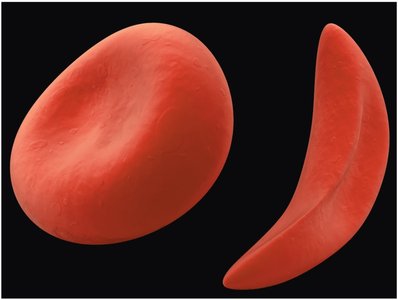
Sickle Cell Anemia: A Molecular Example
Sickle cell anemia is caused by a single point mutation in the β-globin gene, resulting in the substitution of valine for glutamic acid at the sixth position of the polypeptide chain. This alters hemoglobin structure and causes red blood cells to assume a sickle shape.
From Polypeptide to Protein
Polypeptides vs. Proteins
Polypeptides are linear chains of amino acids synthesized during translation. Once released from the ribosome, polypeptides fold into specific three-dimensional structures to become functional proteins.
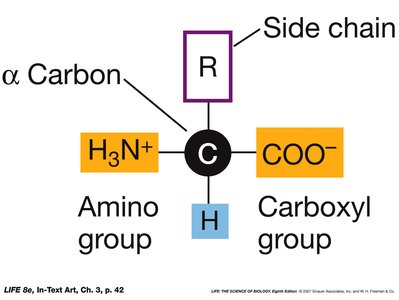
Amino Acid Structure
All amino acids share a common structure: a central (α) carbon bonded to an amino group, a carboxyl group, a hydrogen atom, and a variable side chain (R group). The side chain determines the chemical properties of each amino acid.
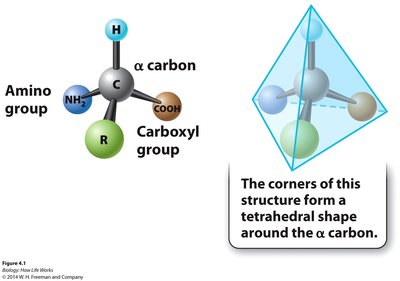
Peptide Bond Formation
Amino acids are linked by peptide bonds, forming the primary structure of proteins. The sequence of amino acids determines the protein's final structure and function.
Levels of Protein Structure
Primary Structure
The primary structure is the unique linear sequence of amino acids in a polypeptide chain, encoded directly by the gene.
Example: The UNC-73/Trio protein contains 2463 amino acids in its primary structure.

Secondary Structure
Secondary structure refers to local folding patterns within a polypeptide, stabilized by hydrogen bonds. The two main types are the α-helix and β-pleated sheet.
α-Helix: A right-handed coil stabilized by hydrogen bonds between every fourth amino acid.
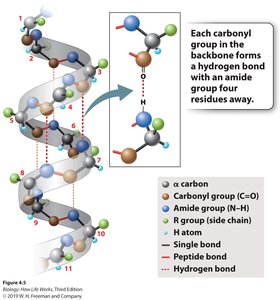
β-Pleated Sheet: Formed by hydrogen bonding between parallel or antiparallel segments of the polypeptide chain.
Tertiary Structure
The tertiary structure is the overall three-dimensional shape of a single polypeptide chain, determined by interactions among side chains (R groups) and the peptide backbone. This structure is essential for protein function and can be dynamic.
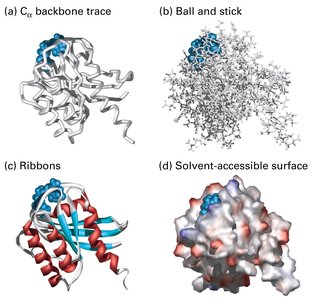
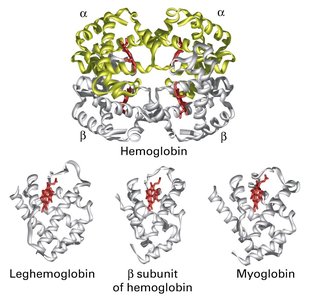
Quaternary Structure
Quaternary structure arises when two or more polypeptide chains (subunits) assemble into a functional protein complex. Hemoglobin is a classic example, consisting of two alpha and two beta chains.
Summary Table: Levels of Protein Structure
Level | Description | Stabilizing Forces |
|---|---|---|
Primary | Linear sequence of amino acids | Peptide bonds |
Secondary | Local folding (α-helix, β-sheet) | Hydrogen bonds |
Tertiary | 3D structure of a single polypeptide | Hydrogen bonds, ionic bonds, hydrophobic interactions, disulfide bridges |
Quaternary | Assembly of multiple polypeptides | Same as tertiary, plus subunit interactions |
