 Back
Back(W3 L1) Translation: Mechanisms and Comparison in Prokaryotes and Eukaryotes
Study Guide - Smart Notes

Translation
Overview of Translation
Translation is the process by which ribosomes synthesize proteins using messenger RNA (mRNA) as a template. It is a fundamental step in gene expression, occurring in the cytoplasm of both prokaryotic and eukaryotic cells. Translation follows transcription, where DNA is transcribed into mRNA.
Key Requirements: mRNA, ribosomes, tRNA, amino acids, aminoacyl-tRNA synthetase, GTP, initiation/elongation/release factors, post-translational modifications.
Role of Aminoacyl-tRNA Synthetase: Catalyzes the attachment of amino acids to their corresponding tRNAs, forming aminoacyl-tRNA complexes ready for translation. This process requires ATP.
Prokaryotic Translation
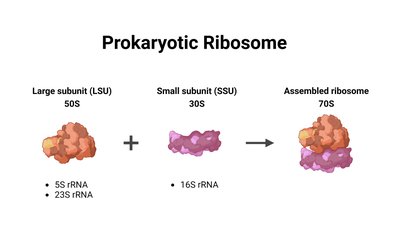
Structure of the Prokaryotic Ribosome
The prokaryotic ribosome is a 70S complex, composed of a 30S small subunit and a 50S large subunit. The small subunit contains 16S rRNA and 21 proteins, while the large subunit contains 23S and 5S rRNA and 34 proteins. The ribosome is responsible for mRNA binding, decoding, and peptide bond formation.
Small Subunit (30S): Binds mRNA and decodes codons.
Large Subunit (50S): Catalyzes peptide bond formation.
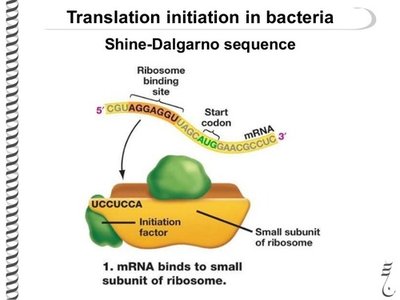
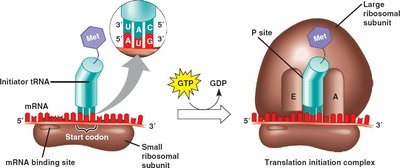
Initiation of Translation in Prokaryotes
Initiation positions the ribosome on the mRNA so the start codon is in the correct place and the initiator tRNA can begin protein synthesis. The Shine-Dalgarno sequence, a purine-rich region upstream of the start codon, aligns the ribosome for translation by base-pairing with the 16S rRNA.
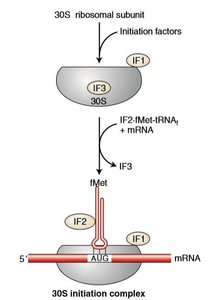
Initiation Factors: IF1, IF2, and IF3 assist in assembling the ribosome and positioning the initiator tRNA.
Initiator tRNA: In bacteria, tRNAfMet carries N-formylmethionine (fMet).
Formation of 70S Initiation Complex: The 50S subunit joins after correct codon-anticodon pairing, forming the complete ribosome ready for elongation.
Elongation in Prokaryotes
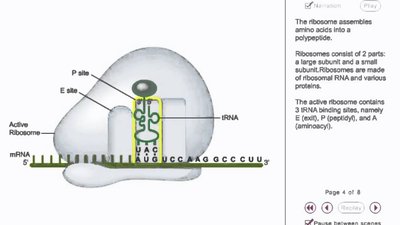
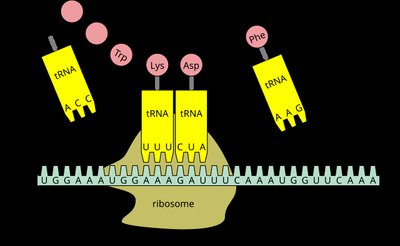
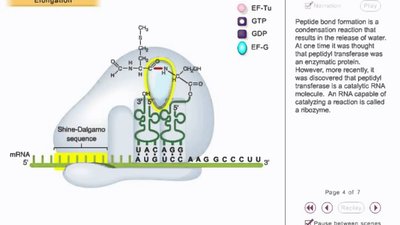
Elongation is the repeating cycle that adds amino acids to the growing polypeptide chain as the ribosome moves along the mRNA. It involves the A (aminoacyl), P (peptidyl), and E (exit) sites, and elongation factors EF-Tu, EF-Ts, and EF-G.
Codon Recognition: Aminoacyl-tRNA binds to the A site via codon-anticodon pairing, assisted by EF-Tu and GTP.
Peptide Bond Formation: Peptidyl transferase catalyzes the bond between the amino acid in the A site and the polypeptide in the P site.
Translocation: The ribosome moves one codon forward, shifting tRNAs and making space for the next aminoacyl-tRNA.
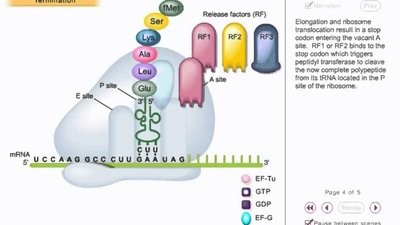
Termination in Prokaryotes
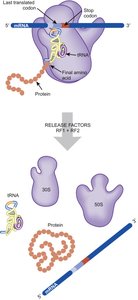
Termination occurs when a stop codon enters the A site, signaling the release of the polypeptide and recycling of the ribosome. Release factors RF1, RF2, and RF3 recognize stop codons and promote hydrolysis of the polypeptide from the tRNA in the P site.
Stop Codons: UAA, UAG, UGA.
Release Factors: RF1 (UAA, UAG), RF2 (UAA, UGA), RF3 (removal of RF1/RF2).
Ribosome Recycling: RRF and EF-G split the ribosome into subunits for reuse.
Post-Translational Modifications: Folding, phosphorylation, glycosylation.
Eukaryotic Translation
Structure of the Eukaryotic Ribosome
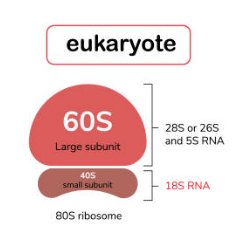
The eukaryotic ribosome is an 80S complex, composed of a 40S small subunit and a 60S large subunit. The small subunit contains 18S rRNA, while the large subunit contains 28S (or 26S), 5S rRNA.
Small Subunit (40S): Binds mRNA and decodes codons.
Large Subunit (60S): Catalyzes peptide bond formation.
Initiation of Translation in Eukaryotes
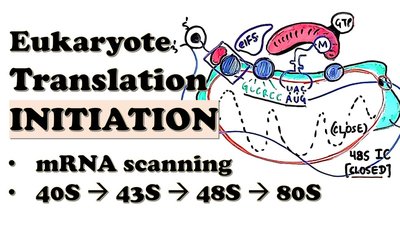
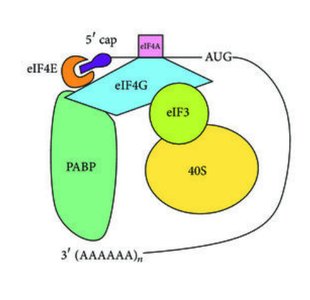
Initiation recruits the ribosome to the mRNA, finds the correct start codon, and assembles an 80S ribosome with the initiator tRNA in the P site. Most eukaryotic mRNAs have a 5′ cap and a 3′ poly(A) tail, recognized by initiation factors.
eIF4F Complex: eIF4E binds the cap, eIF4G links factors, eIF4A unwinds secondary structure, PABP binds poly(A) tail.
43S Pre-Initiation Complex: 40S subunit binds initiation factors and Met–tRNAi.
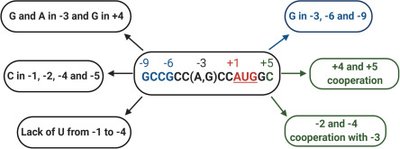
Scanning and Kozak Sequence: 40S scans the 5′ UTR to find the AUG start codon in a Kozak consensus sequence.
Formation of 80S Initiation Complex: 60S subunit joins after correct start codon recognition.
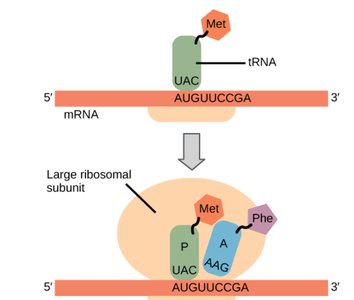

Elongation in Eukaryotes
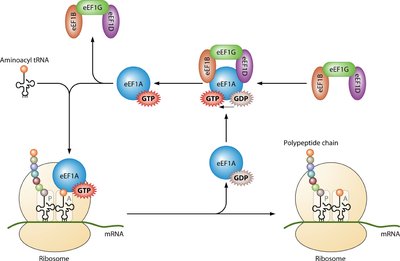
Elongation in eukaryotes is a repeating cycle that adds amino acids to the growing polypeptide chain as the 80S ribosome moves along the mRNA. It relies on elongation factors eEF1A, eEF1B, and eEF2, and the ribosome’s A, P, and E sites.
Codon Recognition: Aminoacyl-tRNA binds to the A site via codon-anticodon pairing, assisted by eEF1A and GTP.
Peptide Bond Formation: Peptidyl transferase catalyzes the bond between the amino acid in the A site and the polypeptide in the P site.
Translocation: The ribosome moves one codon forward, shifting tRNAs and making space for the next aminoacyl-tRNA.
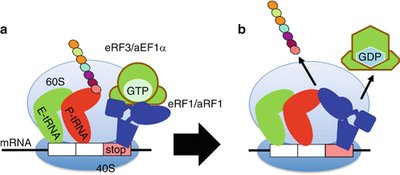
Termination in Eukaryotes
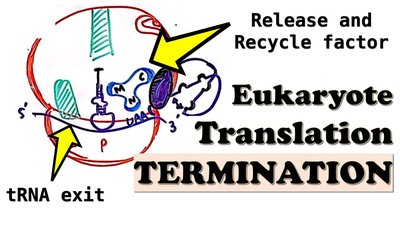
Termination occurs when a stop codon enters the A site, triggering release of the completed polypeptide and recycling of the ribosome. Eukaryotic release factors eRF1 and eRF3 recognize all three stop codons and promote hydrolysis of the polypeptide from the tRNA in the P site.
Stop Codons: UAA, UAG, UGA.
Release Factors: eRF1 (all stop codons), eRF3 (GTPase partner).
Ribosome Recycling: ABCE1 splits the ribosome into subunits for reuse.
Post-Translational Modifications: Folding, phosphorylation, glycosylation.

Translation Summary
Comparison of Translation in Prokaryotes and Eukaryotes
The following table summarizes the key differences between translation in prokaryotes and eukaryotes:
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Ribosome Size | 70S (30S + 50S) | 80S (40S + 60S) |
Initiation Site | Shine-Dalgarno sequence | Kozak sequence |
Initiation Factors | IF-1, IF-2, IF-3 | eIFs |
First tRNA | Formyl-methionine (fMet) | Methionine (Met) |
Elongation Factors | EF-Tu, EF-G | eEF1, eEF2 |
Termination Factors | RF-1, RF-2, RF-3 | eRF1, eRF3 |
Location | Cytoplasm | Cytoplasm (RER for secretory proteins) |
Transcription-Translation Coupling | Yes | No (Occurs separately) |
DNA Replication, Transcription, and Translation Summary
Comparison of the Three Processes
The table below compares DNA replication, transcription, and translation:
Feature | DNA Replication | Transcription | Translation |
|---|---|---|---|
Template | DNA | DNA | mRNA |
Enzymes Involved | DNA polymerase | RNA polymerase | Ribosome, aminoacyl-tRNA synthetase |
Product | Two identical DNA molecules | mRNA | Protein |
Energy Requirement | ATP, dNTP hydrolysis | NTP hydrolysis | GTP hydrolysis |
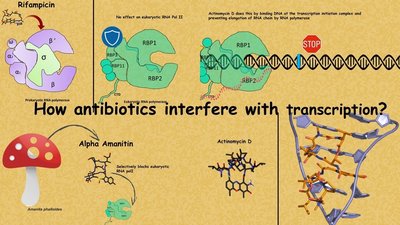
Case Study: Antibiotics Targeting Bacterial Protein Synthesis
Antibiotics and Protein Synthesis
Understanding the differences in translation and transcription between prokaryotes and eukaryotes has enabled the development of antibiotics that selectively target bacterial protein synthesis. For example, tetracyclines and aminoglycosides bind bacterial ribosomes, preventing translation, while macrolides and lincosamides disrupt bacterial translation machinery. Quinolones inhibit bacterial DNA gyrase and topoisomerase IV, essential for DNA replication.
Example: Antibiotics such as rifampicin, alpha-amanitin, and actinomycin D interfere with transcription by targeting bacterial RNA polymerase or DNA, selectively inhibiting bacterial processes without affecting eukaryotic cells.

Additional info: The notes above expand on brief points and provide academic context for each step, including definitions, examples, and comparisons. All images included are directly relevant to the explanation of the adjacent paragraph.
