 Back
BackTranslation: Molecular Mechanisms and Components
Study Guide - Smart Notes

Translation: Molecular Mechanisms and Components
Flow of Genetic Information
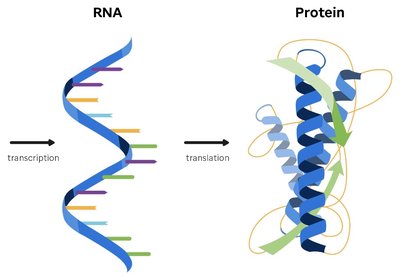
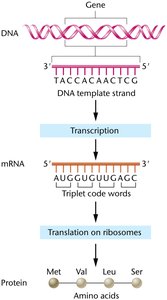
The flow of genetic information in cells follows the central dogma: DNA is transcribed into RNA, which is then translated into protein. This process ensures that genetic instructions are converted into functional molecules.
DNA: Stores genetic information in the nucleus.
mRNA: Acts as an unstable intermediate, transferring information from the nucleus to the cytosol.
Protein Synthesis: Uses the genetic code to convert nucleotide sequences into proteins.
Genetic Code: The set of rules by which nucleotide sequences are translated into amino acid sequences.
tRNA: Structure and Function
Transfer RNAs (tRNAs) are essential adapters that translate mRNA codons into amino acids during protein synthesis. Their structure and modifications are crucial for accurate translation.
Function: tRNAs carry specific amino acids to the ribosome, matching mRNA codons via their anticodon region.
Anticodon: A triplet sequence in tRNA that pairs with the complementary codon in mRNA.
Polypeptide Attachment: The growing polypeptide remains attached to tRNA during translation.
Transcription: tRNAs are transcribed as large precursors by RNA Polymerase III in eukaryotes and then processed into mature forms.
Length: Mature tRNAs are 75-90 ribonucleotides long.
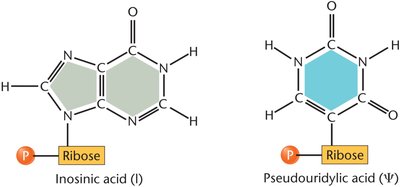
Modified Bases: tRNAs contain unique bases such as inosinic acid and pseudouridylic acid not found in mRNA.
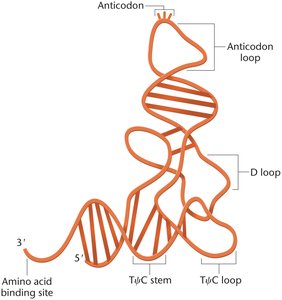

tRNA Features and the Wobble Hypothesis
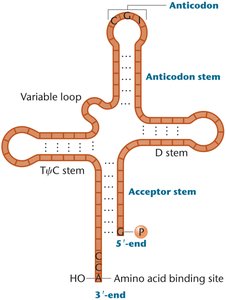
tRNAs fold into a cloverleaf stem-loop structure, with modified bases concentrated in loop regions and the wobble position. The wobble hypothesis explains how tRNAs can recognize multiple codons, increasing translation efficiency.
Cloverleaf Structure: Formed by base-pairing within the tRNA molecule.
Anticodon Loop: Contains the anticodon sequence.
3'-CCA End: Site of amino acid attachment.
Wobble Position: The third base of the codon is less constrained, allowing nonstandard base pairing.
Minimum tRNA Species: About 30 tRNAs can accommodate 61 amino acid codons due to wobble pairing.
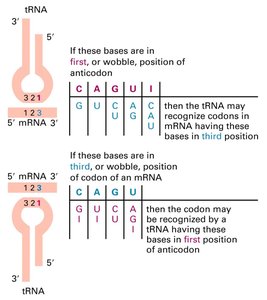
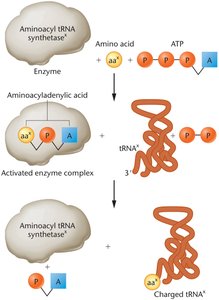
Charging tRNA Molecules
Charging refers to the covalent attachment of an amino acid to its corresponding tRNA, catalyzed by aminoacyl-tRNA synthetases. This process is essential for translation fidelity.
Enzymes: 20 different aminoacyl-tRNA synthetases, one for each amino acid.
Steps:
Amino acid reacts with ATP to form aminoacyladenylic acid (activated form).
The complex binds to the specific tRNA, attaching the amino acid.
The charged tRNA is released and ready for translation.
Translation: Initiation, Elongation, and Termination
Translation is the process by which ribosomes synthesize proteins using mRNA as a template. It is divided into three stages: initiation, elongation, and termination, each requiring specific factors and ribosomal components.
Initiation: mRNA associates with the small ribosomal subunit, requiring charged tRNA, GTP, Mg2+, and initiation factors (IF proteins). In bacteria, the Shine-Dalgarno sequence aligns the ribosome with the start codon.
Elongation: The ribosome forms P and A sites for tRNA binding. Peptide bonds are catalyzed by rRNA in the large subunit. Elongation requires GTP and elongation factors (EFs).
Termination: Release factors (RFs) recognize stop codons, catalyzing the release of the polypeptide and dissociation of the ribosomal complex.
Protein Factors in E. coli Translation
Specific protein factors facilitate each stage of translation in E. coli:
Process | Factor | Role |
|---|---|---|
Initiation | IF1 | Binds 30S subunit, prevents premature tRNA binding to A site |
Initiation | IF2 | Binds initiator fMet-tRNA, transfers to P site, releases upon GTP hydrolysis |
Initiation | IF3 | Prevents premature association of 30S and 50S subunits |
Elongation | EF-Tu | Binds GTP, brings aminoacyl-tRNA to A site |
Elongation | EF-Ts | Regulates EF-Tu activity |
Elongation | EF-G | Stimulates translocation, GTP-dependent |
Termination | RF1 | Releases polypeptide, specific for UAA and UAG codons |
Termination | RF2 | Releases polypeptide, specific for UGA and UAA codons |
Termination | RF3 | Stimulates RF1 and RF2 release |
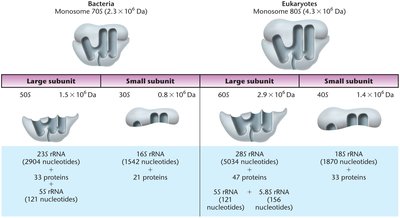
Ribosome Structure and Function
Ribosomes are large cellular structures composed of proteins and rRNAs. They are the site of protein synthesis and are divided into large and small subunits, named by their Svedberg coefficients.
Prokaryotic Ribosomes: 70S, composed of 50S (large) and 30S (small) subunits.
Eukaryotic Ribosomes: 80S, composed of 60S (large) and 40S (small) subunits.
rRNAs: Catalytic components of ribosomes.
Subunit Separation: Ribosome splits into subunits when not actively synthesizing protein.

Translation Initiation in Bacteria
Initiation in bacteria involves the Shine-Dalgarno sequence, which aligns the mRNA start codon with the ribosomal P site. IF2 plays a direct role in stabilizing the mRNA and initiator tRNA at the P site, and the large subunit joins last to complete initiation.
Translation Elongation in Bacteria
Elongation involves the sequential addition of amino acids to the growing polypeptide chain:
Charged tRNA enters the A site.
Peptide bond formation and translocation occur.
Uncharged tRNA exits via the E site.
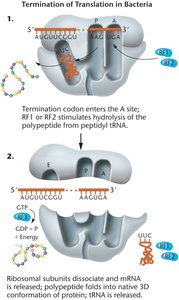
Translation Termination in Bacteria
Termination occurs when a stop codon is recognized by RF1 or RF2, leading to the release of the polypeptide and dissociation of the ribosomal complex.

Summary Table: Ribosome Composition
Bacteria | Eukaryotes | |
|---|---|---|
Monosome | 70S | 80S |
Large Subunit | 50S | 60S |
Small Subunit | 30S | 40S |
rRNA Components | 23S, 5S, 16S | 28S, 5.8S, 5S, 18S |
Protein Components | 33 (large), 21 (small) | 49 (large), 33 (small) |
Key Equations
Peptide Bond Formation:
tRNA Charging:
Additional info:
tRNA modifications (inosinic acid, pseudouridylic acid) enhance stability and decoding flexibility.
Wobble base pairing allows a single tRNA to recognize multiple codons, reducing the number of tRNAs required.
Ribosome subunit composition and Svedberg coefficients are critical for distinguishing prokaryotic and eukaryotic translation machinery.
