 Back
BackTransmission Genetics: Mendelian Principles, Probability, and Pedigree Analysis
Study Guide - Smart Notes

Transmission Genetics
Gregor Mendel and the Foundations of Genetic Transmission
Gregor Mendel's experiments with pea plants established the basic principles of heredity, which underpin modern genetics. His systematic approach and use of the scientific method led to the discovery of predictable patterns in the inheritance of traits.
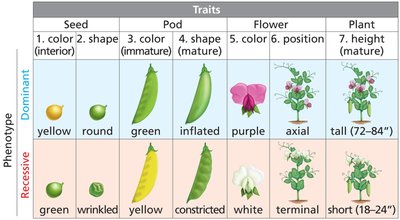
Mendel's Background: Mendel studied natural sciences and chose peas for their clear, dichotomous traits.
Blending Theory vs. Particulate Inheritance: Mendel disproved the blending theory, which posited that offspring traits were intermediate mixtures of parental traits, by demonstrating the reappearance of parental traits in subsequent generations.
Scientific Method: Mendel's approach included observation, hypothesis formation, controlled experimentation, data collection, and interpretation.
Experimental Innovations and Life Cycle of Pea Plants
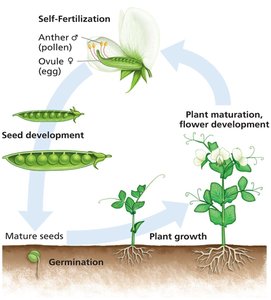
Mendel's success was due to several critical innovations in his experimental design and understanding of the pea plant life cycle.
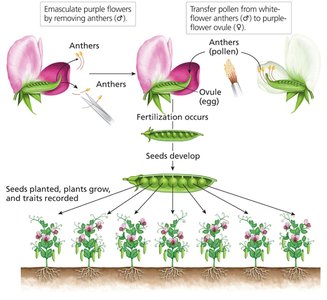
Controlled Crosses: Mendel controlled pollination to ensure accurate parentage.
Pure-Breeding Strains: He established strains that consistently produced the same phenotype.
Dichotomous Traits: Traits with only two possible phenotypes were selected.
Quantification: Mendel counted and analyzed large numbers of offspring.
Replicate, Reciprocal, and Test Crosses: These allowed for robust analysis of inheritance patterns.
Monohybrid Crosses and Segregation of Alleles
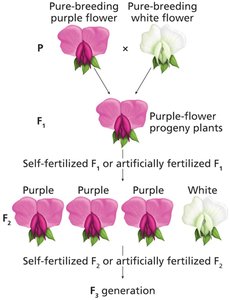
Monohybrid crosses involve the inheritance of a single trait. Mendel's experiments revealed the concepts of dominance, segregation, and predictable ratios in offspring.
Homozygous vs. Heterozygous: Pure-breeding parents are homozygous; F1 offspring are heterozygous.
Dominance: One phenotype dominates in the F1 generation.
Segregation: The recessive phenotype reappears in the F2 generation.
Ratios: F2 generation shows a 3:1 phenotypic ratio and a 1:2:1 genotypic ratio.
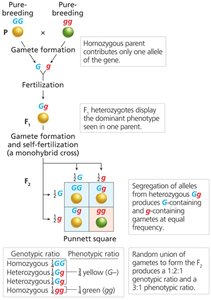
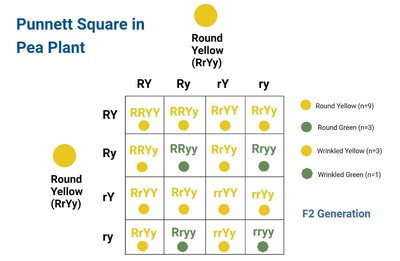
Mendel's First Law: Law of Segregation
The law of segregation states that each individual has two alleles for each trait, which segregate during gamete formation, resulting in predictable inheritance patterns.
Punnett Square: A tool to visualize allele segregation and offspring ratios.
Alleles: Hereditary units that determine phenotype.
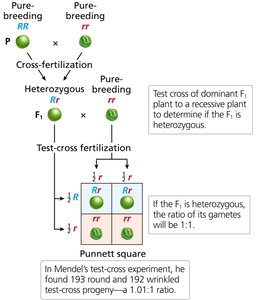
Test-Cross Analysis
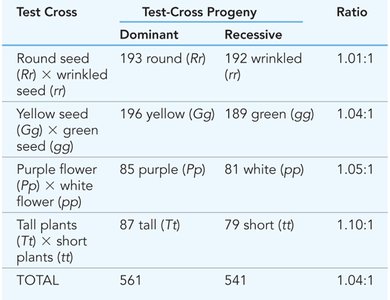
Test crosses are used to determine the genotype of an individual by crossing it with a homozygous recessive partner.
1:1 Ratio: If the tested individual is heterozygous, offspring will show a 1:1 ratio of dominant to recessive phenotypes.
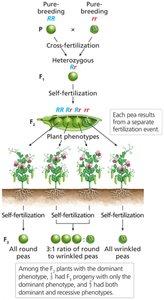
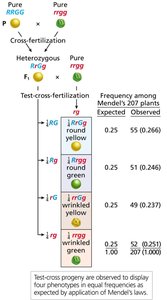
Genotype Determination via F3 Progeny
Self-fertilization of F2 plants and analysis of F3 progeny can reveal the genotype of F2 individuals.
F3 Analysis: F2 plants with only dominant phenotype produce only dominant F3 progeny; those with both phenotypes are heterozygous.
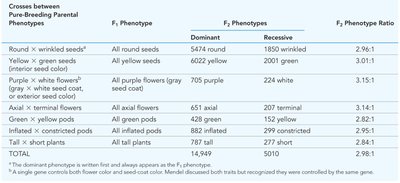
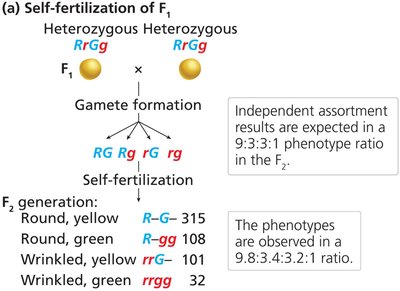
Dihybrid and Trihybrid Crosses: Law of Independent Assortment
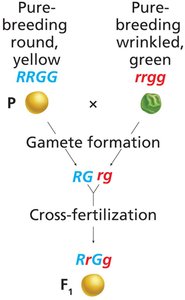
Mendel extended his analysis to crosses involving two or more traits, leading to the law of independent assortment.
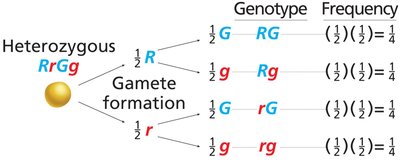
Independent Assortment: Alleles of different genes segregate independently during gamete formation.
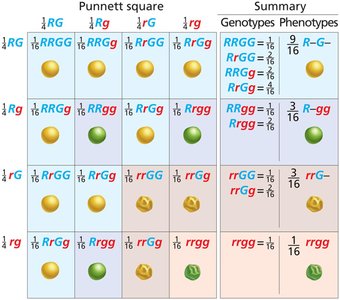
9:3:3:1 Ratio: Dihybrid crosses produce a characteristic phenotypic ratio in the F2 generation.
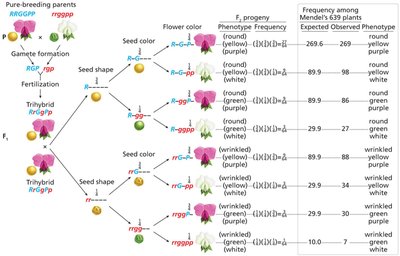
Trihybrid Crosses: Used to further test independent assortment.
Probability Theory in Genetics
Probability rules are essential for predicting genetic outcomes. Mendel recognized the role of chance in inheritance.
Product Rule: Probability of independent events occurring together is the product of their individual probabilities.
Sum Rule: Probability of mutually exclusive events is the sum of their individual probabilities.
Conditional Probability: Probability modified by prior information about the outcome.
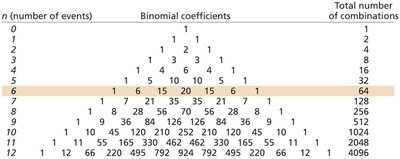
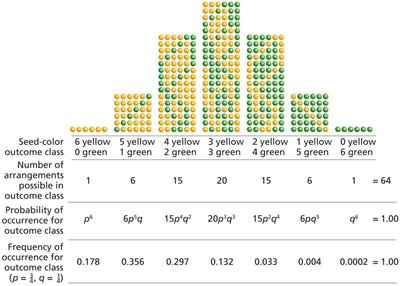
Binomial Probability: Used for predicting the likelihood of combinations of outcomes in a series of events.

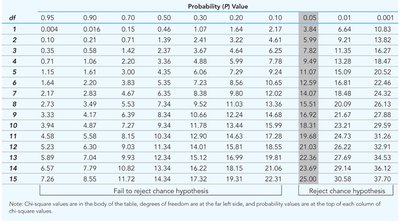
Chi-Square Goodness of Fit
The chi-square test quantifies how closely observed genetic outcomes match expected ratios, providing a statistical basis for evaluating hypotheses.
Formula: , where O = observed, E = expected.
P Value: Probability that observed deviations are due to chance.
Degrees of Freedom: Number of outcome classes minus one.
Statistical Significance: P < 0.05 indicates rejection of the hypothesis of chance.

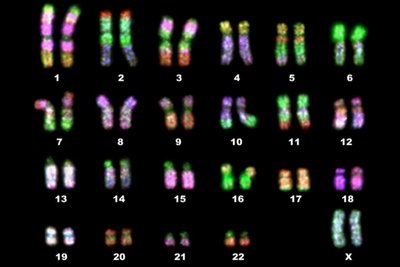
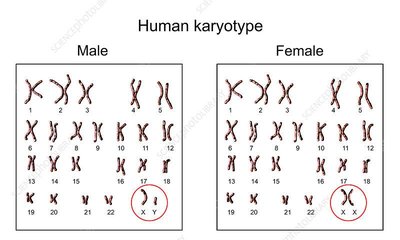
Pedigree Analysis and Autosomal Inheritance
Pedigrees are used to trace inheritance patterns in humans and animals, applying Mendelian principles to autosomal traits.
Autosomal Inheritance: Genes located on autosomes (non-sex chromosomes) are inherited equally by males and females.
Pedigree Symbols: Standard notation is used to represent individuals, relationships, and trait status.

Autosomal Dominant and Recessive Inheritance
Autosomal traits can be dominant or recessive, each with characteristic patterns in pedigrees.
Autosomal Dominant: Trait appears in every generation; affected individuals have at least one affected parent.
Autosomal Recessive: Trait often skips generations; affected individuals may have unaffected, heterozygous parents.
Molecular Genetics of Mendel’s Traits
Modern molecular genetics has identified the DNA sequence differences underlying Mendel’s classic traits, linking transmission genetics to molecular function.
Seed Shape (Sbe1): Dominant allele produces starch-branching enzyme; mutant allele results in wrinkled seeds.
Stem Length (Le): Dominant allele produces gibberellin; mutant allele leads to short plants.
Seed Color (Sgr): Dominant allele enables chlorophyll breakdown; mutant allele causes green seeds.
Flower Color (bHLH): Dominant allele activates anthocyanin production; mutant allele results in white flowers.
Summary Table: Molecular Identification of Mendel’s Traits
The following table summarizes the molecular basis of four Mendelian traits:
Trait | Gene and Product | Dominant Allele Function | Mutant Allele Function |
|---|---|---|---|
Seed shape | Sbe1 (starch-branching enzyme) | Converts amylose to amylopectin | Loss of enzyme, wrinkled seeds |
Stem length | Le (gibberellin 3β-hydroxylase) | Produces gibberellin, tall plants | Low gibberellin, short plants |
Seed color | Sgr (chlorophyll breakdown enzyme) | Breaks down chlorophyll, yellow seeds | No breakdown, green seeds |
Flower color | bHLH (transcription factor) | Activates anthocyanin production, purple flowers | No activation, white flowers |
Central Conclusions from Molecular Studies
Inheritance of alleles parallels phenotypic transmission.
Phenotypic variation results from differences in protein structure and function.
Molecular analysis identifies DNA sequence differences and their consequences.
Functional analysis of protein products explains phenotypic outcomes.
