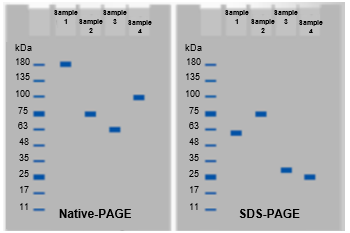
SDS PAGE, or Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis, is a powerful technique used to separate proteins based on their mass. Unlike native PAGE, which maintains the proteins' native structures and interactions, SDS PAGE denatures proteins, allowing for the analysis of individual subunits. This method is particularly useful for separating protein subunits that are not covalently linked. However, when covalent bonds, such as disulfide bonds, are present, additional steps are necessary to fully separate these subunits.
Beta mercaptoethanol (often abbreviated as βME) is a reducing agent that cleaves disulfide bonds, making it essential for analyzing proteins with quaternary structures. For instance, consider a protein composed of four subunits: A, B, C, and D. In this scenario, subunits A and C are linked by a disulfide bond, while subunits B and D are connected only through non-covalent interactions. When subjected to native PAGE, the entire protein complex appears as a single band due to the retention of its native properties, influenced by the shape, mass, and charge of the protein.
In contrast, when SDS is introduced, it denatures the proteins and imparts a negative charge, allowing them to migrate through the gel based solely on size. However, SDS does not break disulfide bonds, meaning that subunits A and C remain associated and travel together as a single band. This band will be the largest due to the combined mass of subunits A and C, while subunits B and D will separate based on their individual sizes, resulting in distinct bands for each.
To fully separate all subunits, βME is added prior to SDS PAGE. This treatment cleaves the disulfide bond between subunits A and C, allowing them to migrate independently. Consequently, the largest band will now represent only subunit A, followed by subunit B, and then subunit C, with subunit D remaining the smallest and fastest migrating band. This strategic use of βME in conjunction with SDS PAGE provides valuable insights into protein structure and interactions, enabling a clearer understanding of the individual components of complex proteins.